Académique Documents
Professionnel Documents
Culture Documents
1 s2.0 S014181301530088X Main PDF
Transféré par
a3kt95Titre original
Copyright
Formats disponibles
Partager ce document
Partager ou intégrer le document
Avez-vous trouvé ce document utile ?
Ce contenu est-il inapproprié ?
Signaler ce documentDroits d'auteur :
Formats disponibles
1 s2.0 S014181301530088X Main PDF
Transféré par
a3kt95Droits d'auteur :
Formats disponibles
International Journal of Biological Macromolecules 82 (2016) 10411054
Contents lists available at ScienceDirect
International Journal of Biological Macromolecules
journal homepage: www.elsevier.com/locate/ijbiomac
Review
Classication, mode of action and production strategy of xylanase and
its application for biofuel production from water hyacinth
Uma Shankar Prasad Uday a , Payel Choudhury b , Tarun Kanti Bandyopadhyay a ,
Biswanath Bhunia c,
a
Department of Chemical Engineering, National Institute of Technology, Agartala 799046, India
Department of Integrated Energy studies, National Institute of Technology, Agartala 799046, India
c
Department of Bio Engineering, National Institute of Technology, Agartala 799046, India
b
a r t i c l e
i n f o
Article history:
Received 17 September 2015
Received in revised form 26 October 2015
Accepted 27 October 2015
Available online 1 November 2015
Keywords:
Xylanase
Lignocellulose
Water hyacinth
Recombinant DNA technology
Biofuel
a b s t r a c t
Xylanases are classied under glycoside hydrolase families which represent one of the largest groups of
commercial enzymes. Depolymerizing xylan molecules into monomeric pentose units involves the synergistic action of mainly two key enzymes which are endo--xylanase and -xylosidase. Xylanases are
different with respect to their mode of action, substrate specicities, biochemical properties, 3D structure
and are widely produced by a spectrum of bacteria and fungi. Currently, large scale production of xylanase
can be produced through the application of genetic engineering tool which allow fast identication of
novel xylanase genes and their genetic variations makes it an ideal enzymes. Due to depletion of fossil fuel, there is urgent need to nd out environment friendly and sustainable energy sources. Therefore,
utilisation of cheap lignocellulosic materials along with proper optimisation of process is most important
for cost efcient ethanol production. Among, various types of lignocellulosic substances, water hyacinth,
a noxious aquatic weed, has been found in many tropical. Therefore, the technological development for
biofuel production from water hyacinth is becoming commercially worthwhile. In this review, the classication and mode of action of xylanase including genetic regulation and strategy for robust xylanase
production have been critically discussed from recent reports. In addition various strategies for cost effective biofuel production from water hyacinth including chimeric proteins design has also been critically
evaluated.
2015 Elsevier B.V. All rights reserved.
Contents
1.
2.
3.
Introduction . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1042
Xylanase: its classication and mode of action . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1043
2.1.
Family 5 glycoside hydrolase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1043
2.2.
Family 8 glycoside hydrolase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1043
2.3.
Family 10 glycoside hydrolase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1043
2.4.
Family 11 glycoside hydrolase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1044
2.5.
Family 19 glycoside hydrolase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1044
2.6.
Family 30 glycoside hydrolase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1044
Strategy for xylanase production . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1045
3.1.
Strategy with wild producer . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1045
3.2.
Strategy for recombinant producer . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1045
3.2.1.
Cloning of xylanase gene for enhanced xylanase expression . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1045
3.2.2.
Mutagenesis for enhanced xylanase expression . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1045
Corresponding author.
E-mail address: bbhunia@gmail.com (B. Bhunia).
http://dx.doi.org/10.1016/j.ijbiomac.2015.10.086
0141-8130/ 2015 Elsevier B.V. All rights reserved.
1042
4.
5.
6.
7.
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
Genetic regulation of xylanase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1047
Fermentative production of xylanase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1047
5.1.
Optimisation of medium parameters . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1048
5.1.1.
Xylanase production through submerged fermentation (SmF) . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1048
5.1.2.
Xylanase production through solid state fermentation (SSF) . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1048
5.1.3.
Media design by statistical approaches . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1049
5.2.
Optimisation of physicochemical parameters design . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1049
Strategy to improve cost effective ethanol production from water hyacinth using xylanase . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1050
Conclusion . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1051
Acknowledgment . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1052
References . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 1052
1. Introduction
Biotechnology is one important scientic eld where the highest development during the last few decades has been taken place.
It is the integration of biological, physical and engineering sciences in order to achieve technological application using biological
systems [1,2]. Microorganisms such as bacteria, yeasts and lamentous fungi are used in most biotechnological processes, however
vascular plants, algae and even animal tissue can also be utilised
[3]. Recent application of biotechnology has been focused on biological products such as enzymes, antibiotics, hormones etc. One
of the advantages of the advanced biotechnology process over
classical biotechnology is that the mass balance for biotechnological products and control mechanism can easily evaluated [3,4].
Nevertheless, there have been very rapid application of genetic
engineering in bioprocess leading to optimisation of the production of established or novel metabolites of commercial importance
[2,4].
Enzymes are used to produce a wide range of biotechnological
products utilised by the food, chemical, and allied industries and
have been already recognised as valuable catalysts for production
of ne chemicals and pharmaceuticals and several organic transformations [2,5,6]. The estimated value of world enzyme market
was about US $3060 billion in 2010 [7]. Table 1 summarises the
major producers of industrial enzymes. The majority of the industrial enzyme production belongs to Novo Industri A/S (Denmark),
Danisco/Genencor (Denmark and USA), BASF (Germany) and DSM
(Netherlands), controlling about 73% of the market between them
[2,8]. It has been found that most of the enzymes industrially produced so far function in a narrow range of temperature, pH and ionic
strength which are prerequisite for the commercial application of
these enzymes [9,10]. Hence, the search for a new enzyme having wide working range of those function parameters is a primary
requirement [7].
Hydrolase represents one of the largest groups of industrial
enzymes and accounts for about 75% of total worldwide enzyme
sales. The carbo-hydrolases, signify one of the second largest
groups of industrial enzymes which can be produced by bacteria,
fungi, animal and plant cells [11]. Carbo-hydrolases include cellulases and hemicellulase which act on cellulose and hemicellulose
which are two major fundamental components of lignocellulose
generated through photosynthesis in plant. These two carbohydrate polymers are covalent cross-linked with a non-carbohydrate
Table 1
Major producers of industrial enzymes [2].
Company
Market share (%)
Novo Industri A/S (Denmark)
Danisco/Genencor (Denmark and USA)
BASF (Germany)
DSM (Netherlands)
Others
45
17
4
5
29
polymer lignin [12,13]. The cellulose is a principal framework of the
cell which consists of chains of glucose linked by -1,4 linkages. The
strong hydrogen bonds play an important role to form the cellulose
chains into micro-brils, leads to crystalline in nature and very
recalcitrant to degradation. However, some parts of the cellulose
is prone to degrade whose structure may be amorphous in nature.
The cellulose is further embedded in a matrix of hemicellulose [14].
Hemicellulose is a branched hetero-polymer consisting of pentose (d-xylose and d-arabinose) and hexose (d-mannose, d-glucose
and d-galactose) sugars where xylose is most abundant. Hemicellulose is the one of the most abundant second renewable biomass
in nature after cellulose, which accounts for 2535% of lignocellulosic biomass [15]. The most abundant hemicellulose in nature
is xylan which contains mainly -d-xylopyranosyl residues linked
by -1,4-glycosidic bonds [15,16]. In plants, an overlying layer of
xylan and cellulose forms through hydrogen bonding which is further covalently linked with lignin leads to formation of an outside
sheath for the protection of the plant. Xylan accounts for 3035% of
total dry weight, however, the exact amount of the xylan will differ
from plants to plants which display a large variation in composition
during extraction from different sources [16,17].
India is an agricultural country; hence agro-industrial wastes
and by-products, weeds etc. are in abundance here which includes
wheat bran, sugar cane bagasse, corn cobs, rice bran and water
hyacinth etc. Agricultural and forest industries produce ca 40 million tonnes of lignocellulose wastes which contain approx. 3040%
of hemicellulose in dry weight [18]. Due to improper handling of
these waste materials it is a source of environmental pollution. The
processing units which are generating agro-waste materials/byproducts are struggling hard for their conversion into value added
products. Therefore, they are taking interest to utilise even a little bit of resources that can play a signicant role in the economic
uplift of a country. Therefore, utilisation of neglected materials to
produce value added products can be a challenge for all researchers
which can be further employed in different industries. Enzyme,
carbo-hydrolase is the prominent option for production from these
wastes that contain high amount of cellulose and hemicellulose.
Production of cellulase and its applications have been investigated
deeply by several researchers; however, production and application of xylanase are still in infant stage. Xylanases is used widely
in different industries for its wide applications the most common among them is in production of biofuel, majority chemicals,
animal feeds to release pentose sugars by enzymatic treatment,
bio-bleaching of wood pulps, food additives in baking industry,
ingredients in laundry detergents or fabric care compositions [19].
As wastes are inexpensive and easily processed, the production of
xylanase using manipulation of biotechnological techniques with
its wide application can offer an economical solution to the problem of environment pollution in near future. Therefore, this review
addresses the recent progress in classication and mode of action,
genetic regulation, strategy of robust xylanase production and its
industrial applications.
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
1043
2. Xylanase: its classication and mode of action
Due to the structural heterogeneity of hemicellulose, complete degradation requires the synergistic action of different
hemicellulase enzymes. These enzymes are endo-xylanase (endo1,4--xylanase, E.C.3.2.1.8), -xylosidase (xylan-1,4--xylosidase,
E.C.3.2.1.37), -glucuronidase (-glucosiduronase, E.C.3.2.1.139),
-arabinofuranosidase (-l-arabinofuranosidase, E.C.3.2.1.55) and
acetylxylan esterase (E.C.3.1.1.72), among the endoxylanases and
-xylosidases (collectively xylanases) are the two major enzymes
which are responsible for the hydrolysis of xylan. Endo-xylanases
act on homopolymeric backbone of 1,4-linked -d-xylopyranose
for production of xylooligomers and -xylosidases act on these
xylooligomers to release xylose [20,21].
According to the CAZy database (http://www.cazy.org),
xylanases (EC 3.2.1.8) are classied under glycoside hydrolase
(GH) families 5, 7, 8, 9, 10, 11, 12, 16, 26, 30, 43, 44, 51 and 62.
Nevertheless, the orders of classied in families 16, 51 and 62
are bi-functional enzymes as it contains two catalytic domains
which are unlike families i.e., 5, 7, 8, 10, 11 and 43, they have a
truly distinct catalytic domain with endo-1,4--xylanase activity
[22]. The remaining families 9, 12, 26, 30 and 44 are also found,
which may have residual or secondary xylanase activity. However,
most scientic classication has been carried out based on the
hydrophobic cluster analysis of the catalytic domains and similarities in the amino acid sequences and xylanases have been primarily
classied as GH 10 and 11 [23]. Although recently the members of
these two families have been thoroughly explored, the catalytic
properties of the members of the remaining families (5, 7, 8 and
43) it has are recent and the information remains very limited
[24]. It has been found that the members of GH families 5, 7, 8,
10, 11 and 43 are different with respect to their physicochemical
properties, structure, mode of action and substrate specicities
[22]. The salient feature of GH 10 family is a high molecular
mass structurally composed of a cellulose-binding domain and a
catalytic domain connected by a linker peptide with a pI between
8 and 9.5. This family normally has a (/) 8 fold TIM barrel.
GH 11 family (or family G) has a low molecular mass and lower
pI values. Based on their isoelectric points (pI), GH 11 family is
further sub-grouped into two, alkaline and acidic pIs [20,25,26].
Fig. 1. Biological assembly image for 3GTN (crystal structure of XynC from Bacillus subtilis 168. Protein chains are coloured from the N-terminal to the C-terminal
using a rainbow (spectral) colour gradient) (http://www.rcsb.org/pdb/home/home.
do). (For interpretation of the references to colour in this gure legend, the reader
is referred to the web version of this article.)
Fig. 2. Biological assembly image for 2B4F (structure of A cold-adapted family 8
xylanase in complex with substrate. Protein chains are coloured from the N-terminal
to the C-terminal using a rainbow (spectral) colour gradient) (http://www.rcsb.org/
pdb/home/home.do). (For interpretation of the references to colour in this gure
legend, the reader is referred to the web version of this article.)
2.1. Family 5 glycoside hydrolase
Though there is very limited information regarding the family
of 5 glycoside hydrolase GH 5), however, a distinctive endo mode
of action of this family has been investigated. It has the capability
of hydrolysing the beta-1,4 xylan chain at a specic site directed by
the position of an alpha-1,2-linked glucuronate moiety. Recently,
xylanase C (XynC, kD 90.86) from Bacillus subtilis 168, is under the
GH 5 xylanase has been cloned (Fig. 1). The gene is over-expressed
and crystallised. It has been found from crystallography study that
this family allows the exceptional unique specicity and the role
for biological depolymerisation and processing of glucuronoxylan
[27].
2.2. Family 8 glycoside hydrolase
This family also has capability to hydrolyse the beta-1,4 xylan
chain. The ligand binding follows induced t mechanism and during
catalytic events a number of conformational changes occur with a
repositioning of the proton donor into a more catalytically competent position. pXyl (GH-8 xylanase, Fig. 2) from the Antarctic
bacterium Pseudoalteromonas haloplanktis TAH3a has been studied
to investigate the mode of action of this family. Therefore the X-ray
crystallography study has been carried out in complex of enzyme
(pXyl, kD 47.34) with its substrate xylopentaose and product
xylotriose. It has been found that the subsites (3 to +3) are responsible to give a depth insight into the structurefunction relationship
of family 8 glycoside hydrolases. Furthermore, the mechanism of
action of GH-8 members is described mentioning that the substrate
hydrolysis is preceded by a conformational change without the
substrate ground-state chair conformation [28].
2.3. Family 10 glycoside hydrolase
Santos and his colleagues (2010) had investigated that the
molecular basis for the action mode of TpXyl10B (kD 81.62) from
hyper thermophilic bacterium Thermotoga petrophila RKU-1 at high
temperatures are used for biochemical characterisation and crystallographic methods in the native state and in complex with
xylobiose (Fig. 3). It has been found that there are two subunits
responsible for bonding with substrate. The xylanase (TpXyl10B)
showed a temperature-dependent action mode due to coupling
effect of temperature-induced structural changes. The temperature dependency action mode was further conrmed by molecular
dynamics simulations method suggesting that the catalytic loop
(Trp297-Lys326) is responsible for signicant modications in the
product release area, which drives the enzymatic activity to the
specic release of xylobiose at high temperatures [29].
1044
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
Fig. 3. Biological assembly image for 3NIY (crystal structure of native xylanase 10B
from Thermotoga petrophila RKU-1. Protein chains are coloured from the N-terminal
to the C-terminal using a rainbow (spectral) colour gradient) (http://www.rcsb.org/
pdb/home/home.do). (For interpretation of the references to colour in this gure
legend, the reader is referred to the web version of this article.)
2.4. Family 11 glycoside hydrolase
The pH optima of this family are distinguished by the nature
of the residue adjacent to the acid/base catalyst. Bacillus circulans
xylanase (BCX, kD 20.7) contains an asparagine residue at position
35 and the activity depends on pH which is due to ionisation of
the nucleophile Glu78 (pKa 4.6) and the Glu172 (pKa 6.7) (Fig. 4).
The pH optima were shifted from 5.7 to 4.6 and approximately 20%
enhancement of enzyme activity was found due to substitution of
asparagine residue in place of aspartic acid residue (N35D BCX). It is
obvious that the apparent pKa values of N35D BCX were in the range
of 3.55.8 which may be due to 3.7 (Asp35), 5.7 (Glu78) and 8.4
(Glu172). It was reported that both enzymes (BCX and N35D BCX)
follow a double-displacement mechanism with Glu78 which acts
as nucleophile and Asp35 and Glu172 function together as the general acid/base catalyst. In case of N35D BCX, the enzyme showed a
reverse protonation mechanism and showed activity when Asp35
and Glu78 are protonated and de-protonated respectively which
indicates mutant enzyme has higher catalytic efciency than wild
type. It has been revealed that the increase in efciency of N35D
BCX and all acidic family 11 xylanases, is due to the creation of
a short (2.7A) hydrogen bond between Asp35 and Glu172 that will
ultimately stabilise the transition state for glycosyl transfer [30].
Fig. 4. Biological assembly image for 1C5H (hydrogen bonding and catalysis: an
unexpected explanation for how a single amino acid substitution can change the ph
optimum of a glycosidase. Protein chains are coloured from the N-terminal to the
C-terminal using a rainbow (spectral) colour gradient) (http://www.rcsb.org/pdb/
home/home.do). (For interpretation of the references to colour in this gure legend,
the reader is referred to the web version of this article.)
Fig. 5. Biological assembly image for 4HU8 (crystal structure of a bacterial Ig-like
domain containing GH 10 xylanase from termite gut. Protein chains are coloured
from the N-terminal to the C-terminal using a rainbow (spectral) colour gradient)
(http://www.rcsb.org/pdb/home/home.do). (For interpretation of the references to
colour in this gure legend, the reader is referred to the web version of this article.)
2.5. Family 19 glycoside hydrolase
Han et al. (2013) inspected the biochemical characterisation
and crystal structure of a bacterial xylanase, Xyl-ORF19 (kD
41.18) which has been derived from gut bacteria found in woodfeeding termite (Globitermes brachycerastes). It was reported that
xylanase, Xyl-ORF19 has two domains, includes a C-terminal GH
10 catalytic domain and an N-terminal bacterial Ig-like (Big 2)
non-catalytic domain (Fig. 5). The catalytic domain has a (/)8
barrel as it presents in most GH 10 xylanases, along with two
extra beta-strands. The non-catalytic domain is mimic to an
immunoglobulin-like domain of intimins. In terms of kinetic
parameters, pH and temperature proles it has been found that
the enzyme shows fairly low catalytic activity without the noncatalytic domain, which suggested the non-catalytic domain can
affect the biochemical enzyme and biophysical properties [31].
2.6. Family 30 glycoside hydrolase
Glucurono-xylanase Xyn30D (kD 44), a modular enzyme under
family 30 glycoside hydrolase contains a catalytic domain along
with carbohydrate binding module of the CBM35 family (Fig. 6).
Fig. 6. Biological assembly image for 4QB1 (structure of CBM35 from Paenibacillus
barcinonensis protein chains are coloured from the N-terminal to the C-terminal
using a rainbow (spectral) colour gradient) (http://www.rcsb.org/pdb/home/home.
do). (For interpretation of the references to colour in this gure legend, the reader
is referred to the web version of this article.)
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
This family contains a catalytic domain which folds into an (/)8
barrel like as family 10 xylanase with an associated -structure.
The CBM35 includes -sandwich that contains two calcium ions.
However, these two domains fold in an independent manner. The
linker region between two domains allows a moderate exibility for folding of two domains through polar interactions with the
catalytic domain. The ancillary Xyn30D-CBM35 domain was only
shown binding ability with glucuronic acid-containing ligands due
to presence of two aromatic residues allocated a wider pocket. In
this pocket, region E was named where a non-conserved Glu129
makes a link with calcium. The binding ability of Xyn30D-CBM35
to different xylans was proposed due to orientation of negative
charge that extends up to binding pocket leads to interaction with
substrate [32].
3. Strategy for xylanase production
The xylanase enzymes are produced by a wide range of microorganisms including bacteria and fungi, plant and animal cell [2022].
They are either extracellular or intracellular in nature. However, the
interest in microbial xylanase increased to meet the current crisis
of energy demand in the world along with the inability of the plant
and animal xylanase. It has been found that there are two strategies
applied to date for microbial xylanase production. The enzyme produced is either using wild microorganism or recombinant microbial
strain.
3.1. Strategy with wild producer
There is extensive research being carried out for xylanase production from wild type of fungi and bacteria owing to their potential
industrial applications. Screening for proper microorganism is
an important criterion in order to get the desired product. The
microorganism which will be used for xylanase production should
produce adequate yields and should not produce toxins or any
other undesired products. The main challenges with the wild strain
include the availability of potent microbial strain and wide application of this biocatalyst for industrial scale or large scale production.
It is also important to note that this strain should be robust under
industrial conditions. It has been found that there is no batch to
batch variation of xylanase production found using wild strain.
However, low yield is the main drawback for xylanase production using wild strain. Additionally, isolation of potent strain for
production of desired product is tedious and time consuming.
The microbial diversity during degradation of hemicellulosic
material under natural conditions needs evaluation and isolation, screening and characterisation of new well-adapted microbial
strains are used for potentially improved enzyme-production.
There has also been substantial progress for production of xylanase
using wild microbial system such as bacteria; Burkholderia sp. [33]
and Bacillus pumilus [34,35], Actinomycetes; S. actuosus [36], S. cyaneus [37], S. halstedii [38], S. matensis [39], S. olivaceoviridis [40,41],
S. roseiscleroticus [42], S. thermoviolaceus [43] and S. viridosporus
[44] and fungal species; Aspergillus niger [45], P. thermophila [46],
T. longibrachiatum [47] and T. languginosus [48].
Although, a large number of bacterial and fungal species are
also reported as potent source of xylanase, however, lamentous
fungi are the most promising candidate for commercial production of xylanase as they secrete much higher amounts of xylanase
enzymes into the medium. It has been found that xylanases are
mostly produced by mesophilic and thermophilic microorganism
[12]. The mesophilic fungi under the genera Aspergillus and Trichoderma are potent producer for xylanase. Study of thermophilic
microorganisms as potential xylanase producers has recently been
initiated as they have highest temperature optima among microbes
1045
for their growth. Therefore, it is expected that produced enzymes
would have greater stability which can widen the application of
xylanase. In recent years, there is an intensive research for isolation of thermophilic microorganisms such as C. thermophile, H.
insolens, H. lanuginosa, H. grisea, Melanocarpus albomyces, P. variotii,
T. byssochlamydoides, T. emersonii, T. lanuginosus and T. aurantiacus
[12,4951].
3.2. Strategy for recombinant producer
The main challenges for the engineered strategy include the
availability of tools which can be modied by recombinant DNA
technology and the application of these tools so that a desired
xylanase will be produced with high yield and robustness under
industrial conditions. In this strategy, the enzyme which is more
suitable for industrial applications has been chosen [16]. The main
challenge for recombinant DNA technology is to improve the
fermentation characteristics of genetically engineered organisms
by introducing genes. It has been found that the robustness of
engineered xylanase enzymes is often required for industrial applications. There are several reports where cloning of xylanase gene,
random mutagenesis, site specic mutagenesis, or the combination of both have been frequently used to get robust engineered
xylanase enzymes for industrial applications [52]. Iterative saturation mutagenesis (ISM) is a directed evolution method to improve
the favourable characteristics of enzymes. The repetitive cycles of
saturation mutagenesis are applied in ISM at chosen sites of two
or three amino acids of the protein and protein structure. Benecial mutations were found by performing 34 rounds of ISM and
these benecial mutations are systematically incorporated into the
libraries [53]. Few recent research reports on engineering xylanase
are discussed here.
3.2.1. Cloning of xylanase gene for enhanced xylanase expression
Microorganisms secrete different types of xylanase in presence
of different nutritional components. In addition, microorganisms
have ability to develop diverse mechanisms in the induction of
xylanase genes in response to various inducers. Therefore, the
microbial regulatory network is the most advanced approach and
cloning of specic gene is considered to improve productivity
and specicity of xylanase production. Driss et al. (2012) experimented with over production on xylanase GH 11(PoXyn2) from
Penicillium occitanis Pol6. cDNA of PoXyn2 having 320 amino acids
that had been sub-cloned into the pGAPZA vector to construct
recombinant xylanase. The six histidine residues were attached
at the N-terminal. They were further integrated into the genome
of Pichia pastoris X-33. The glyceraldehyde 3-phosphate dehydrogenase (GAP) constitutive promoter was used as control. The
expression of PoXyn2 in P. pastoris was conrmed through activity
assay and SDS-PAGE analysis. The his-tagged helped to purify this
enzyme through one step purication using afnity chromatography (Ni-NTA resin). The over expression of PoXyn2 was reported
with the specic activity of 8549.85 U mg1 in presence of Oat Spelt
Xylan. One step purication strategy helps to decrease the production cost which will widen the applicability of xylanase [54]. A new
xylanase (xyn186) from Alternaria sp. HB186 was cloned to study
the over expression of xylanase. It was found that sequence contains a 748 bp open reading frame which is separated by one intron
having the size of 52 bp. The xyn186, cDNA was prepared by DpnImediated intron deletion which was cloned into pHBM905A and
transformed into P. pastoris GS115 where the gene copy number
was evaluated by the Real-time PCR [55].
3.2.2. Mutagenesis for enhanced xylanase expression
Xylanases are commonly used in many industrial processes such
as pulp bleaching, baking, detergent and biofuels production. It is
1046
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
well known that the characteristics of industrial enzyme should
have longer acting catalytic activity, amenable to novel substrates,
or more stability in presence of chemical environment which is
required to expand their applicability [56]. The above such characteristics can be improved by several strategies such as rational and
non-rational mutagenesis, including random mutagenesis, sitespecic mutagenesis and site-saturation mutagenesis [57].
3.2.2.1. Mutagenesis for enhanced thermostable xylanase expression.
The rational approach includes computational analysis to design a
mutagenesis library in targeted regions which can identify specic
important residues which were further randomly mutagenised. It
is reported that the thermostability of GH 10 xylanase from A. niger
was improved by mutagenesis. The rational approach was applied
through computational analysis to design of a mutagenesis library
in single domain GH 10 xylanase, Xyn10A ASPNG which contains
thermal important residues. The single domain was randomly
mutagenised through iterative saturation mutagenesis (ISM). It
has been found that ve residues (R25W/V29A/I31L/L43F/T58I)
showed 30 folds higher thermal half-life (t1/2 ) at 60 C than wildtype enzyme. In addition, the melting temperature (Tm ) of mutants
was also increased by 17.4 C vs the wild type. It is reported that
N-terminal coil of the enzyme is very mush important for stabilisation of GH 10 xylanase. It has been found that the hydrophobic
interactions between N- and C-terminal ends and within Nterminal elements are responsible to improve the thermostability
of Xyn10A ASPNG [58].
Sometimes it has been found that combination of distinct
hemicellulases and auxiliary enzymes, mainly endo-xylanases and
-xylosidases, increase the biotechnological applications. Therefore several researchers design bi-functional enzyme which can
increase the accessibility of substrate. A bi-functional enzyme
was prepared with fusion of GH 11 endo-1,4--xylanase and GH
43 -xylosidase, both from B. subtilis. According to quaternary
arrangement and accessibility to the substrate, the design was carried out. This chimaera enzyme along with parental enzymes was
successfully expressed in Escherichia coli and the enzyme was puried and characterised. It is interested to note that 3-fold xylose
conversion of fusion enzyme was found in presence of beechwood
xylan and hemicelluloses from pre-treated biomass. In addition
to it, the chimeric enzyme is found which have higher thermalresistance along with a optimum temperature can shift at higher
range i.e., from 35 to 50 C for xylosidase activity. From circular
dichroism unfolding studies, it is understood that stability of the
-xylosidase domain is responsible for improvement of thermal
stability [59].
The thermal-stability of family xylanase AuXyn11A, GH 11 from
Aspergillus usamii E001, was improved by N-terminus replacement
which was constructed by substituting the N-terminal 33 amino
acids of AuXyn11A with the corresponding 38 ones of EvXyn11TS,
a hyper thermostable G11 family xylanase. Rational computational
method was applied to prepared hybrid xylanase, named AEXynM.
The hybrid along with parental gene was expressed in P. pastoris
GS115. The specic activities of two recombinant xylanases were
found to be 10,437 and 9529 U mg1 respectively. The temperature optimum was shifted for hybrid xylanase vs parental from
70 to 75 C and stability of hybrid xylanase was much higher
than parental xylanase. The melting temperature (Tm ) was analysed by the Protein Thermal Shift (PTS) method and found to
increase by 34.0 C as compared with that of parental enzyme.
AExynM xylanase was further mutagenised by site-directed mutagenesis and three single mutant genes from AExynM (AExynMC5T,
AExynMP9S, and AExynMH14N) were constructed which were further expressed in P. pastoris GS115. It is interesting to note that the
thermo-stabilities of three recombinant mutants clearly decreased
as compared with AExynM xylanase which is due to fact that the
three amino acids (Cys-5, Pro-9, and His-14) is responsible for thermal stability which was replaced in mutant [60].
3.2.2.2. Mutagenesis for enhanced pH stable xylanase expression. The
alkaline stability of the xylanase from T. lanuginosus was enhanced
by directed evolution using error-prone PCR mutagenesis. The positive clone which produced clear zones on pH 9 and 12 of xylan agar
plates were screened. It was found that the variant NC38 can withstand at pH 10 retaining 84% activity for 90 min at 60 C which is
higher than parental enzyme. The variant NC38 was further cloned
into pBGP1 under the control of GAP to promote and pET22b(+)
expression in P. pastoris and Escherichia coli BL21, respectively. A
higher extracellular expression of xylanase was found in P. pastoris (261.7 U ml1 ) in comparison with E. coli (47.9 U ml1 ) and
total activity obtained in P. pastoris was 545-fold higher than E. coli.
This alkaline stable mutated xylanase favours the application of this
enzyme in the pulp and paper industry [61].
The pH-dependent activity was also carried out by mutagenesis
and the pH dependency of wild type B. circulans xylanase (BcX) is
due to the pKa values of its nucleophile Glu78 and general acid/base
Glu172. Therefore several strategies were applied by manipulating
these pKa values to nd out the shifting of the pH (opt). It has been
found that random succinylation had no signicant effect on shifting of pH optimum of xylanase activity as it is expected to change
of global charge. Mutation has further carried out residues near
or within the active site of BcX, however, little effect was found
regarding change of pH optimum due to lowering the apparent pKa
value of Glu78.
Substitution of Glu172 with a His resulted lowered the pH (opt)
of BcX from 5.6 to 4.7 with retaining 8% activity towards a xylobioside substrate His lowered the pH (opt) of BcX from 5.6 to 4.7
while retaining 8% activity towards a xylobioside substrate. It was
revealed by NMR spectroscopy that utilizing a reverse protonation mechanism was applied in spite of the opposite charges of
the introduced residues. The reverse protonation mechanism indicates that the pKa value of the general acid is lower than that of the
nucleophile, and therefore, only a small fraction of population of
enzyme is in a catalytically competent ionisation state. However,
it is a fact that overall activity is maintained due to the increased
strength of the general acid [62].
3.2.2.3. Look-Through Mutagenesis (LTMTM ) and Combinatorial Benecial Mutagenesis (CBMTM ). Recently Look-Through Mutagenesis
(LTMTM ) and Combinatorial Benecial Mutagenesis (CBMTM ) are
commonly used in mutagenesis. LTM is used for rapid screening of
amino acids at selected positions in the protein sequence which can
introduce favourable properties whereas CBM is a method to identify the best assembly of individual mutations. These techniques are
advantageous as they do not required high-throughput screening or
special equipment. These techniques are designed to get the maximum information through the small, chemistry-driven libraries
which lead to identify informative leads. Still, there are several
reports which found that several rational and random mutagenesis
strategies have been reported to improve the favourable properties of xylanase. However, Look-Through Mutagenesis (LTM) and
Combinatorial Benecial Mutagenesis (CBM) are more reliable on
structural data, which are not always found to effectively search
the sequence space using random/saturation mutagenesis. Iterative saturation mutagenesis (ISM) focuses on small regions of the
protein which is determined from examining statistical analyses
of sequence/function/stability relationships and protein structure
while LTM/CBM scans the whole protein. LTM/CBM would most
likely be used to nd out the benecial mutations which are present
throughout the protein. Both methods incorporate data into design
libraries with reduced numbers of variants which will lower the
cost of the screening effort [63]. LTM/CBM was applied to nd
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
1047
Fig. 7. Cellular recognition, genetic regulation and expression of xylanase.
out the thermostable mutants of xylanase from Hypocrea jecorina (GenBank Accession No. P36217), having the 32-residue signal
sequence. The LTM xylanase libraries were prepared by polymerase
cycling assembly (PCA) and further cloned through expression vector pPICZC into P. pastoris. Each oligonucleotide contributes to a
21 base pair homology with its reverse-orientated neighbouring
oligonucleotide. The complete xylanase gene was assembled with
16 oligonucleotides, containing an average length of 63 nucleotides.
Ten selected benecial xylanase mutations were obtained through
the LTM screens to improve thermo-tolerance. These mutants were
used to combinatorially build a CBM library of 360 mutants by the
use of degenerate oligonucleotides. These oligonucleotides were
further designed to limit the number of non-consensus amino acids
created by the degenerate oligonucleotides. CBM library was produced by PCR by the degenerate oligonucleotides. The library was
further sub-cloned into the pPICZC plasmid and transformed into
the P. pastoris X-33 strain and six hundred clones were analysed
for higher temperature optimum for xylanase activity. A diverse
set of novel mutations including N71D, Y73G, T95G and Y96Q were
discovered. It is interesting to note that when a single construct
(Hjx-81) was made with these mutations and the puried protein
retained its activity even after heating at 100 C for 20 min. Therefore, LTM/CBM method is a time-effective method which should
be used for quickly improving the favourable properties of other
industrial and therapeutic enzymes [63].
4. Genetic regulation of xylanase
There are several papers published on cellular recognition of
xylan and genetic regulation and expression of xylanase enzyme
in presence of such complex carbohydrate. It has been found that
xylanases are synthesised when microorganisms are cultured on
xylan since it has induced the activity of enzyme complexes in
microorganisms [64]. It is interesting to note that xylan did not
directly enter inside the cell in order to inuence the regulation of
gene and expression of xylanases enzyme. Since enzyme secretion
is an induction process, there should be a physical contact between
part of the regulatory machinery of the cell and the inducer. The
inducers have some recognition site on the surface of cell which
regulates the process. The expressed enzyme will secrete extracellularly and hydrolyse the polymer to oligosaccharides which can
easily be transported inside the cell with the help of -xyloside
permeases (Fig. 7). These oligosaccharides will trigger the expression of the xylanolytic genes depending upon their concentration.
It has been reported that in the presence of the xylanolytic inducers
the permease activity of the cells increases [12].
The regulation of xylanase production by Aspergillus spp. has
been discussed in this review. The three xylanases and one
-xylosidase, encoded by the xlnA, xlnB, xlnC and xlnD genes,
respectively are produced by Aspergillus spp. in presence of xylose
or xylan as a carbon sources [65]. However, these genes are activated at the transcriptional level by the transcriptional factor XlnR
[66] and repressed by carbon catabolite repression (CCR) which
is mediated by CreA. CreA is a transcription repressor which can
bind CreAbinding sites which were found in Aspergillus spp. [67].
Aspergillus nidulans creA encodes a protein of 415 amino acids which
contains several features characteristic for DNA binding proteins.
Characteristics of these proteins includes zinc ngers, an alaninerich region and frequently appearing SPXX and TPXX motifs [68].
It has been suggested that the regulation of xylanolytic enzyme is
creA-dependent and mediated which is considered as a double lock
mechanism. It is interesting to note that this mechanism is similar
to the ethanol regulation in Aspergillus sp [69]. CreA is a zinc-nger
type transcription factor and two adjacent and divergent CREA
binding sites are required for CreA for in vivo carbon catabolite
repression [70] which is similar with for T. reesei. However, CreAdependent carbon catabolite repression model for T. reesei is only
a single lock mechanism [71].
The protein turnover of proteasome is an important ubiquitin
mediated biological process which is found in eukaryotes. E3 ligases
are an important molecule which is responsible for attaching ubiquitin to protein substrates [72]. The SCF1s include Skp1, Cullin1,
F-box proteins which are the largest family of E3 ligases. In biological process, SCF1 complexes interact with Skp1 which helps E3
ligase to recognise and target its protein substrates [73]. It has been
found that GrrA and FbxA are a F-box proteins found in Aspergillus
spp. which are responsible for several biological functions, such as
cell cycle, transcription, development, cell signalling, and nutrient
sensing etc. [65,74,75].
5. Fermentative production of xylanase
A fermentation process involving microbial cells requires investigation of raw materials, biomass, and how they are treated and
mixed with other ingredients required for cells to grow well. The
medium is sterilised to eliminate all other living microorganisms
and bring together in a large cylindrical vessel, bioreactor or fermenter, typically equipped with agitators, bafes, air spargers, and
various sensing devices for controlling the fermentation conditions.
1048
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
A pure strain of a microorganism is normally make known to the
vessel. The bioreactor supports the natural process by providing
suitable conditions such as optimum temperature, pH, carbon and
nitrogen sources, nutritional salts, vitamins and oxygen (for aerobic organisms), enabling cells to grow and form metabolites and
enzymes. The cells will start to multiply exponentially after a certain period of lag time and reach a maximum cell concentration
as the medium is depleted. Depending on the type of product, the
concentration levels it produces and the purity desired, the fermentation stage might constitute anywhere between 5% and 50%
of the total xed and operating costs of the process. Therefore, optimal design and operation of a bioreactor frequently dominates the
overall technological and economic performance of the process. To
carry out a bioprocess on a large scale, it is necessary to investigate
and develop three principle areas. To obtain a potent biocatalyst
(microorganisms, animal cell, plant cell, or enzyme) along with
medium optimisation is a primary criterion for a fermentation
process. In addition it is required to create the best possible environment for the catalyst to perform by designing the bioreactor
and operating it in the most efcient way. However, efcient downstream processing is required to separate the desired products from
the reaction mixture for cost effective fermentation process.
The optimisation of medium composition is done to maintain a
balance between the various medium components in production
media which is done for commercial practice. Optimisation helps
minimizing the amount of unutilised components at the end of
fermentation. Research efforts have been paying attention mainly
towards evaluating the effect of various carbon, nitrogenous nutrients, metal ion, phosphate and salt as cost-effective substrates on
the yield of enzymes. However, optimisation of environmental and
fermentation parameters such as pH, temperature, aeration, and
agitation is an important part for cost effective bioprocess.
It has been reported that there are no dened media established for the best production of xylanase from different microbial
sources since each organism has its own nutritional requirement
for maximum enzyme production [76]. Microorganisms are capable of utilizing a great variety of carbon and nitrogen sources by
secreting a range of different metabolite into their environment.
Some key intermediates of primary metabolism serve as splitting
points of biosynthetic pathways leading to end products of primary
and secondary metabolism. Secondary metabolism is regulated by
precursors, carbon sources, nitrogen sources, phosphate, trace elements, induction of enzymes of secondary metabolism, catabolic
repression and inhibition, feedback repression and inhibition, and
control by autoregulators [7779]. The effect of carbon and nitrogen sources on secondary metabolism is trained by several factors
including the type of metabolic pathway, the producing organism,
the type and concentration of the sources and whether cultures are
stationary or submerged.
investigated the effect of different parameters for maximum
xylanase production in shake ask. It has been observed that maximum xylanase production was found in presence of xylan as carbon
source and yeast extract as nitrogen source. However, the optimum
pH and temperature were found to be 8 and 28 C respectively for
high yield of xylanase [81]. It is known that several trace elements
are essential for microbial growth because of their involvement as
metalloenzymes or as enzyme activators. In secondary metabolism,
zinc, iron, manganese and pyridoxine are the most important trace
elements [77,8284]. However, the requirement for specic metal
ions depends on the source of enzyme. Potassium phosphate has
been used as a source of phosphate in most studies [82,8588].
This was shown to be responsible for buffering the medium [82].
Furthermore, the supplementation of salts viz., sodium chloride
and phosphate ion was used as inducers for xylanase secretion
are required for bioenergetic and metabolic processes of microbes
such as pH homeostasis and ATP synthesis. Owing to buffering
action, phosphate ions may cause stabilisation of the pH of the fermentation medium (pH homeostasis) which will indirectly favour
xylanase synthesis. Muller et al. (2014) investigated the limitation of pyridoxine on an A. nidulans which produces xylanase B
(XynB) in trickle bed reactors. It was found that A. nidulans is not
able to synthesise pyridoxine which was used to limit microbial
growth and induced to enzyme synthesis. From the experiment it
was observed that due to absence of pyridoxine growth was completely arrested, while xylanase production in culture was found
to be unaltered. Enzyme production has started after 480 h of
continuous fermentation and xylanase production was found to
be 1026 U. The maximum productivity of XynB (21.14 U/g h) was
achieved when pyridoxine was not added to the medium [84].
The SmF was carried out for xylanase production by B. pumilus
strain MK001. The key variables were screened and optimised for
higher production of xylanase. The optimum condition was found
to be pH of 9.0, agitation of 200 rpm, inoculum size 1.25% (v/v) and
inoculum age of 2 h for maximum enzyme production. The carbon
sources were screened and wheat bran showed maximum xylanase
activity (1220.0 IU ml1 ) than other carbon source tested. Among
different inorganic or organic nitrogen sources, a combination of
yeast extract and peptone (0.66% N2 equivalent) is found to produce the maximum (1288.0 IU ml1 ) xylanase titres. Among various
amino acids, vitamins and surfactants, dl- -phenylalanine, niacin,
and polyethylene glycol (PEG)-3330 maximally enhanced xylanase
production by 136.0% (2880.0 IU ml1 ), 79.0% (2190.0 IU ml1 ), and
107% (2536.0 IU ml1 ) respectively. It was reported that the optimal
xylanase production (2886.0 IU ml1 ) was found in 5 L laboratory
fermenter with 8 h reduction of incubation time. However, wholecell immobilisation of B. pumilus strain MK001 showed better
production of xylanase (up to 4000.0 IU ml1 ). The reducing sugar
of 15.5 g/g with saccharication efciency of 20.0% can be achieved
through chemically pretreated Prosopis juliora followed by enzymatic cocktail (xylanase, cellulase and cellobiase) and an ethanol
yield of 0.36 g/g was observed [89].
5.1.1. Xylanase production through submerged fermentation
(SmF)
Xylanases are normally produced through submerged fermentation (SmF) and solid state fermentation (SSF) processes. However,
xylanases are produced worldwide through SmF which allows better control environment during aerobic fermentation process. It
has been found that SmF is normally preferred when the preparations require more puried enzymes [80]. In SmF, puried xylan
is frequently used in bioprocesses for xylanase production. Most
microorganisms can utilise both inorganic and organic forms of
nitrogen which are required to produce amino acids, nucleic
acids, proteins and other cell wall components. Recently, Tallapragada and Venkatesh (2011) isolated A. niger from garden soil and
5.1.2. Xylanase production through solid state fermentation (SSF)
The solid state fermentation (SSF) has some advantages for
enzymes production due to high volumetric productivity, relatively
higher concentration of the enzymes, requirement for inexpensive fermentation equipment, low capital investment and lower
operating cost [90]. This process is normally employed in developing countries like India, because substrates used for xylanase
production are agro-industrial wastes, weeds (water hyacinth) etc.
which are very inexpensive and easily available. There are several reports where substrates used in solid state fermentations are
sugar cane bagasse, wheat bran, rice bran, saw dust, corn cobs,
banana waste, tea waste etc. [91]. The important factors which
affect microbial xylanase synthesis in an SSF system are selection of
5.1. Optimisation of medium parameters
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
a suitable substrate and potent microbes. However, pre-treatment,
particle size, water content and water activity of substrate as well
as relative humidity, type and inoculum size, temperature, cultivation time, maintenance of uniformity in the environment of
SSF system and gaseous atmosphere, i.e., oxygen consumption rate
and carbon dioxide inuence the production rate. Solid state fermentation (SSF) was carried out using A. niger and A. avus using
agro-industrial residues as substrate. From the experiments it was
found that wheat bran was the best carbon source for maximum
xylanase production for both organisms. It was also revealed that
the xylanase production was 18% and 21% higher on wheat bran
with xylan [92]. SSF was carried out for production of xylanase by
A. fumigatus SK1 using untreated oil palm trunk (OPT) as carbon
source. The highest xylanase production was found in presence
of 80% moisture. The fermentation was carried out at initial pH
5.0 with 125 l of OPT as sole carbon and spore source was inoculated with 1 108 spore/g. The xylanase production was found
to be 418.70 U/g substrates [93]. In another experiment, de-oiled
Jatropha curcas seed cake was used as sole carbon source for the
production of xylanase in SSF using S. thermophile which is one
of the abundant by-product of biodiesel industry. The key factors
such as moisture content, initial pH, inoculum, particle size, temperature, carbon supplementation and inducer were optimised and
xylanase production was found to be 1025 U xylanase/g (deoiled
seed cake) [94].
5.1.3. Media design by statistical approaches
In developing a biotechnological industrial process, designing
the fermentation medium is of critical importance. The fermentation medium affects the product concentration and volumetric
productivity [95,96]. It is also important to reduce the cost of the
medium as much as possible, as this may affect the overall process
economics. Medium screening studies are very time consuming and
expensive. This is because the number of possible media combinations that can be tested and the number of fermentation substrates
that are available are also very large. Thus, statistical optimisation approaches have been used to rapidly identify the variables
which are required for maximum production of useful metabolites. Verma and Satyanarayana (2013) clonned a metagenomic
xylanase gene (Mxyl) into shuttle vector pWH1520 and investigated its expression level in B. subtilis. The recombinant strain
was cultured in presence of xylan as a sole carbon source. The
authors have observed that xylanase production had started after
6 h of commencement of fermentation in presence of xylan. During
fermentation the critical variables were identied and optimised
by change one-variable-at-time (COVT) approach for maximum
production of xylanase. The key variables (xylose, inoculum density, incubation time) were further optimised by response surface
methodology (RSM). RSM study enhanced three-fold increase of
xylanase production 119 U mL1 than COVT [97]. Maciel and his
coworkers (2008) investigated xylanase production using A. niger
LPB 326 through solid-state fermentation with sugarcane bagasse
and soybean meal as carbon and nitrogen source respectively. The
effects of key variables were observed and optimised by applying factorial experimental designs. The maximum xylanase activity
was obtained in a medium containing 6.5 g of sugarcane bagasse
and 3.5 g of soybean meal and 85% initial water content. The optimised nutrient salt solution was (in g/L): CuSO4 0.4, KH2 PO4 1.5 and
CoSO4 0.0012. The xylanase production (3099 IU/g of dry matter)
was found after 4 days of fermentation at 30 C [98].
5.2. Optimisation of physicochemical parameters design
Bioreactor operation conditions show diverse effects on product formation in aerobic fermentation processes depending on
inoculum size, oxygen transfer rate, pH and temperature by
1049
inuencing the metabolic pathways and changing the metabolic
uxes [99,100]. The inoculum size has an important effect on
production of metabolites [101]. A close relationship between a
particular morphology and increased process productivity is characteristic of a number of industrially important fermentations
[102,103]. The pattern of expression of enzyme can be modied
by changes in the inoculum size [101,104]. This may be explained
as due to the limitation in other fermentation medium components.
Inoculum quality, in terms of size, type or age, is of prime importance in determining the outcome of fermentations [105108]. The
incubation temperature and pH of the growth medium are important bioprocess parameter that are normally desired to keep both
these variables constant and at their optimal values throughout
the fermentation process. The inuence of temperature and pH on a
bioprocess can be very different, and since the growth process is the
result of many enzymatic processes the inuence of both culture
parameters on the overall bioreaction is quite complex [99,109].
The inuence of temperature on the maximum specic growth rate
of a microorganism is similar to that observed for the activity of
an enzyme. Specic growth rate is progressively increased with
optimum temperature. Whereas beyond that the rapid decrease
of specic growth rate is observed [109,110]. The mechanism of
controlling temperature in enzyme production is not understood
clearly [111]. However, studies by Frankena et al. (1986) showed
temperature and oxygen controlled applications where it shows a
link existed between enzyme synthesis and energy metabolism in
Bacilli [3,112].
The pH culture strongly affects many enzymatic processes and
several species is transported across the cell membrane. Variation
in pH adjusts acid-base equilibria and uxes of various nutrients,
inducers and growth factors between the abiotic and biotic phase
[82]. The inuence of pH on cellular activity is determined by the
sensitivity of the individual enzymes for changing the pH. Enzymes
are normally active only within a denite pH interval and the
total enzymatic activity of the cell is therefore a complex function
with the environmental pH. Microbial cells have a remarkable ability to maintain the intracellular pH at a constant level even with
large variations in the pH of the extracellular medium [110]. This
parameter affects the morphology and the metabolite production
pattern [113,114]. During fermentation; the aeration rate indirectly
indicates the dissolved oxygen level in the fermentation broth. Different dissolved oxygen proles can be obtained through variations
in the aeration rate; variations in the agitation speed of the bioreactor or use of oxygen rich or oxygen decient gas phase [82,115].
The variation in the agitation speed inuences the extent of mixing
in the shake asks or the bioreactor and will also affect the nutrient
availability. Oxygen transfer also shows diverse effects by inuencing metabolic pathways and changing metabolic uxes on product
formation in aerobic fermentation processes.
Biswas et al. (2010) developed a mutant M. albomyces (mutant
IITD3A) and used for xylanase production where a soluble extract
of wheat straw as the sole carbon source. The response surface
methodology (RSM), a statistical optimisation tool was used for
optimisation of process parameters in shake ask. It was revealed
from statistical analysis that the pH is the most important factor
which affects production of xylanase in a greater extent. The experiment was further scaled up in a 14 L bioreactor and xylanase activity
(415 IU mL1 ) was found after 36 h fermentation at pH of 7.8 with
maximum biomass concentration of 3.2 g/L and productivity was
found to be 11,530 IU L1 h1 . The authors also reported that the
fermentation time could be decreased to 24 h for same xylanase
activity by cycling the pH between 7.8 and 8.2 (Fig. 8) and overall productivity was found to be 16,670 IU L1 h1 . Additionally,
fungal morphology also inuences xylanase production like any
other secondary metabolite. It has been found that the fungal pellet (12 mm) showed maximum activity which could be controlled
1050
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
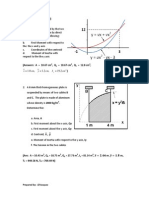
Fig. 8. Effect of aeration rate at 0.25 vvm on production of xylanase by M. albomyces
IITD3A in 14 L bioreactor with pH cycling. (Filled square xylanase activity; lled triangle pH; lled circle dissolved oxygen and inverted lled triangle dry mycelia
mass).
by agitation and pH. Pellets having hairy structure are benecial
for substrate diffusion onto the core of the pellet resulting in high
synthesis of xylanase [116]. As any hyphae growth is not expected
for xylanase production, therefore, the agitation speed was further
increased from 400 to 600 RPM which will force to complete pellet
transformation. This is due to fact that enhanced agitation speed
will cut down the fungal hyphae in small pieces and force to pellet formation. The microscopic observation of hyphae and pellet is
illustrated in Fig. 9. Therefore, the viscosity of fermentation media
is decreased and DO is increased due to pellet formation which
can result in improved mixing and mass transfer phenomenon. The
maximum xylanase activity was improved from 415 to 480 IU mL1
after 24 h due to increase of agitation speed from 400 to 600 RPM.
However, after further increase in the agitation speed to 700 rpm,
xylanase production was found to be decreased to 430 IU mL1 . This
is due to fact that a higher agitation speed creates the high shear
stress which is deleterious for fungal pellet growth. It has been
found that agitation rate and air ow rate both determine the level
of the dissolved oxygen available in fermentation media which is
also crucial for enhancing productivity of xylanase. Therefore, the
aeration rate was further optimised and optimised aeration rate of
0.25 vvm was reported. The xylanase activity and biomass were
found to be 550 IU mL1 and 1.73 g/L respectively with a very high
overall volumetric productivity of 22,000 IU L1 h1 [117].
It has been found that the shake ask experiments are not appropriate for production of recombinant xylanase which is due to fact
that relatively uncontrolled environment in shake ask leads to
unwanted proteolysis and revert to the wild strain. Therefore, a
fed-batch fermentation system may be appropriate for recombinant xylanase production. Punt et al. (2011) reported that a 25 L
fermenter setup may be appropriate for 1 g/L recombinant protein
production using A. niger through fed batch fermentation. A 200 mL
pre-culture of potato dextrose broth containing 1 106 spores
ml1 was inoculated in the fermenter. Media composition at fermenter was glucose (30 g/L), NaNO3 (10 g/L), MgSO4 7H2 O (0.8 g/L),
KH2 PO4 (2 g/L), CaCl2 2H2 O (0.1 g/L), yeast extract (5 g/L), tryptone
(5 g/L). The experiment was carried out at pH 4.5, T = 30 C, airow of
1.2 L/min. DO was required to maintain about 20% through adjustment of agitation speed which was in the range of 400800 rpm.
Pure oxygen was sparged if agitation speed was not sufcient to
maintain DO value of 20%. The feeding has normally started 24 h
after the start of the fermentation at the rate of 5 g/h. The composition of feed medium was glucose (200 g/L), KH2 PO4 (10 g/L), NaNO3
(30 g/L), yeast extract (10 g/L), tryptone (10 g/L). The fed batch fermentation was terminated 4856 h after start of the feeding [118].
6. Strategy to improve cost effective ethanol production
from water hyacinth using xylanase
There is urgent need to nd out environment friendly and
sustainable energy sources due to rapid industrial development.
Therefore, utilisation of lignocellulosic materials can be considered as sustainable biomass for production of renewable biofuels.
The hydrolysis of this biomass is extremely important for the production of biofuel. Therefore, utilisation of cheap source along
with proper optimisation of process is most important for cost
efcient ethanol production. Among, various types of lignocellulosic substances, water hyacinth (Eichhorni acrassipes), a noxious
aquatic weed has been found potentially in many tropical regions
of the world. Water hyacinth is a monocotyledonous fast growing
freshwater aquatic weed. It is under the family of Pontederiaceae.
There are several protocol established in a number of developing countries, like as India for production of biofuel from water
hyacinth. However, cost minimisation and optimisation of all stages
Fig. 9. Effect of agitation rate on fungal morphology during production of xylanase by M. albomyces IITD3A (a) at 400 rpm and (b) at 600 rpm.
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
can improve the applicability of biofuel. In addition, production
of fuels from any waste material is an important contribution in
the self-sustaining society [119]. The bioethanol production from
water hyacinth has three stages. They are pre-treatment, saccharication and fermentation. The pretreatment of biomass is normally
carried out using acid/alkali treatment. This process formed the
unwanted toxic products which turn down the subsequent fermentation process. Xylan, a hemicellulose is the second most abundant
biopolymer found in most of the lignocellulosic material. There
are a variety of enzymes involved for hydrolysis of xylan into its
constituent sugars which is a raw material for subsequent biofuel
production through fermentation. Two enzymes, mainly endo-xylanase and -xylosidase, are important to hydrolyse the xylan
into xylo-oligosaccharides and xylose units. Therefore selection of
proper enzyme/mixture of enzyme or genetic engineering tool may
be the possible method to achieve this goal [120].
It is important to understand that the composition of water
hyacinth, particularly the specic sugars in the hemicellulose fraction can vary from place to place, which will help for selection of
enzyme for its degradation. The accurate analysis of composition
of particular water hyacinth could impact on the overall economy of the process. The enzymes mostly considered for conversion
of water hyacinth to biofuel are cellulases, endo-xylanase and xylosidase. It is obvious that other enzymes may be added after
evaluating the composition of substrate such as endo-mannanase
and -mannosidase for mannose, -arabino-furanosidase for arabinose and pectinases for pectin [121]. Efciency of mixture of
enzyme action on a substrate can be evaluated on the basis of
the conversion to monomer sugars and the percentage of carbohydrates that were converted. Mostly, the yield of reducing sugar
including glucose and xylose etc. is measured and efciency will be
addressed in terms of respective reducing sugar yield. It is obvious
that the ratios of different enzymes are an important parameter
for yield of reducing sugars [121]. It is interesting to note that certain target for respective sugar yield is required for cost effective
bioethanol production. Kristensen et al. (2009) reported that 4%
(w/w) ethanol concentration in reactor was achieved using reducing sugar yield of 8% (w/w) [122]. It was also mentioned that the
energy requirement was less for distillation which leads the process to be more economical. It was found from another report that
the nal 40 g/L ethanol could be produced from 80 g/L initial sugar
yield [123]. The nal ethanol concentration depends on several
factors such as initial sugar loading, type of microbial culture ant
type of biomass etc. [124]. The level of sugar loaded also depends
on which types of fermentation are carried out including the carbohydrate and lignin composition of the substrate. It is obvious,
the optimisation of enzyme ratio, lower substrate levels are maintained for better synergistic actions of enzymes and yield of ethanol
production.
There are several companies such as Novozymes and Genencor
that produce enzyme mixtures containing about 80 enzymes which
are applied in several laboratories for cost and effectiveness of biofuel production [125]. The exact composition of these mixtures is
not known. Qing and Wyman (2011) reported that higher xylanase
activity is required to improve cost effective bioethanol production.
The main limitation of these commercial enzymes is lack of characterisation [126]. Research may be carried out on how the addition
of puried xylanase enzyme with enzyme mixture may improve
the overall process. It is also noted that some also argued about
the presence of non-essential enzymes in the commercial mixtures. Therefore it is suggested that removal of such could increase
specic activity and enzyme cost [121]. In addition, the composition of enzyme mixtures could be improved by modifying the
ratio of different enzymes by the specic characteristics of the substrate [127]. However, the crude mixtures may be preferable to
single enzymes as they contribute to hydrolysis efciency. In case of
1051
puried enzymes, pH and temperature optima of these enzymes
are the key factors for selection of enzyme. In addition, enzymes
should have high thermal and pH stability under the reaction conditions which can improve the applicability and achieve lower costs
[128].
The grafting of the above three enzymes to chimeric enzymes
could be the alternative path to improve the efciency of the bioprocess and cost effective ethanol production. In addition, it also
decreases the required cost of the applied enzymes. It is obvious
that the primary goal is to decrease the process cost of the overall
bioprocess. In the 21st century, the development of bioprocesses
has been focused on enzyme mediated bioprocess which is an
attractive tool to reach the economical and ecological goals. Many
bioprocesses, mainly biofuel production from lignocellulosic material, lignin-degrading enzymes such as laccase may be incorporated
during fusion of chimeric enzymes. Punt et al. (2011) reported that
the production of a chimeric laccase from P. cinnabarinus may be
linked to cellulose which can act on lignin [118]. Ravalason et al.
(2009) were experimenting with chimeric laccase-CBM1 and interestingly the production of the chimeric laccase was noticed. It is
interesting to note that the produced enzyme showed the biochemical properties of the native laccase, however, optimal temperature
was higher (85 C instead of 65 C), and the thermal stability was
markedly decreased which is may be due to the truncation of a small
part of the protein at the linker position [129]. Pham et al. (2010)
also designed a chimeric enzyme for the biofuel sector. They experimented with fusion of mannanases with cellulases and found them
to act synergistically to enhance lignocellulosic biomass degradation. The production of chimeric mannanases was found 130 mg/l
using A. niger. It was reported that all the kinetic and physicochemical properties were close to the native protein. However, thermal
stability was seven times higher at 65 C than native enzyme which
may be due to the fusion of the linker of CBM part [130]. It was also
reported by several researchers that the CBM is required for protective role on the connected enzyme. However, the deletion of the
CBM leads to turn down the thermo-stability and optimal temperature. In addition glycosylation at the turning point region i.e., in
between the catalytic and the substrate-binding domain is found
that it can increase the thermostability of the chimeric protein
[131133].
7. Conclusion
The value added product such as biofuel from waste like water
hyacinth has received much attention globally by the researchers.
The ethanol production from lignocellulosic material includes three
steps. They are pretreatment, saccharication and fermentation.
The objective is the depolymerisation of lignocellulosic biomass
into fermentable sugars. Pretreatment is normally carried out by
either chemical route using acid/base or biological route using
enzymes/microbial cells. However, during chemical treatment formation of furfural and 5-hydroxymethylfurfural which are sugar
degradation products will inhibit microbial growth in the successive fermentation steps. It has been found that no such products are
formed during enzyme treatment as it is usually conducted under
mild conditions. The enzyme treatment methods are normally
carried out using xylanase and cellulase which can be produced
widely by a spectrum of bacteria and fungi. Filamentous fungi have
potential for industrial production of xylanase. It is obvious that
large scale xylanase can be produced through random mutagenesis,
site specic mutagenesis, Iterative Saturation Mutagenesis (ISM),
Look-Through Mutagenesis (LTMTM ) and Combinatorial Benecial
Mutagenesis (CBMTM ) using applications of genetic engineering
tool. It is obvious that chimeric proteins of cellulose and xylanase
may improve enzyme-based industrial bioprocesses by providing
1052
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
promising options for cost effective biofuel production from water
hyacinth.
Acknowledgment
[30]
This research is supported by the NIT Agartala, Ministry of
Human Resource and Development, Government of India.
[31]
References
[1] A. Moser, Bioprocess Technology, Springer-Verlag, New York, USA, 1988.
[2] C. Ratledge, B. Kristiansen, Basic Biotechnology, 3rd ed., Cambridge
University Press India Pvt. Ltd., New Delhi, 2008.
[3] M.L. Shuler, F. Kargi, Bioprocess Engineering: Basic Concepts, Practice Hall of
India Private Limited, New Delhi, 2008.
[4] M. Moo-Young, Y. Chisti, Biochemical engineering in biotechnology, Pure
Appl. Chem. 66 (1994) 117136.
[5] B. Bhunia, B. Basak, T. Mandal, P. Bhattacharya, A. Dey, Effect of pH and
temperature on stability and kinetics of novel extracellular serine alkaline
protease (70 kDa), Int. J. Biol. Macromol. 54 (2013) 18.
[6] B. Bhunia, D. Dutta, S. Chaudhuri, Extracellular alkaline protease from
Bacillus licheniformis NCIM-2042: improving enzyme activity assay and
characterization, Eng. Life Sci. 11 (2011) 207215.
[7] S. Savitha, S. Sadhasivam, K. Swaminathan, F.H. Lin, Fungal protease:
production, purication and compatibility with laundry detergents and
their wash performance, J. Inst. Chem. Eng. 42 (2011) 298304.
[8] B. Jaouadi, S.E. Chaabouni, M. Rhimi, S. Bejar, Biochemical and molecular
characterization of a detergent-stable serine alkaline protease from Bacillus
pumilus CBS with high catalytic efciency, Biochimie 90 (2008) 12911305.
[9] B. Bhunia, B. Basak, A. Dey, A review on production of serine alkaline
protease by Bacillus spp, J. Biochem. Technol. 3 (2012) 448457.
[10] B. Bhunia, K. Behera, A. Baquee, H. Sharma, Optimization of alkaline protease
activity from Bacillus subtilis 2724 by response surface methodology (RSM),
Int. J. Biol. Sci. Eng. 1 (2010) 158169.
[11] M.K. Bhat, Cellulases and related enzymes in biotechnology, Biotechnol.
Adv. 18 (2000) 355383.
[12] M.L. Polizeli, A.C. Rizzatti, R. Monti, H.F. Terenzi, J.A. Jorge, D.S. Amorim,
Xylanases from fungi: properties and industrial applications, Appl.
Microbiol. Biotechnol. 67 (2005) 577591.
[13] R.A. Prade, Xylanases: from biology to biotechnology, Biotechnol. Genet.
Eng. Rev. 13 (1996) 101131.
[14] K.L. Eriksson, H. Bermek, Lignin, lignocellulose, ligninase, Appl. Microbiol.
Ind. (2009) 373384.
[15] B.C. Saha, Hemicellulose bioconversion, J. Ind. Microbiol. Biotechnol. 30
(2003) 279291.
[16] Q.K. Beg, M. Kapoor, L. Mahajan, G.S. Hoondal, Microbial xylanases and their
industrial applications: a review, Appl. Microbiol. Biotechnol. 56 (2001)
326338.
[17] K. Li, P. Azadi, R. Collins, J. Tolan, J.S. Kim, K.L. Eriksson, Relationships
between activities of xylanases and xylan structures, Enzyme Microb.
Technol. 27 (2000) 8994.
[18] . Cano, C. Palet, Xylooligosaccharide recovery from agricultural biomass
waste treatment with enzymatic polymeric membranes and
characterization of products with MALDI-TOF-MS, J. Membr. Sci. 291 (2007)
96105.
[19] N. Kulkarni, A. Shendye, M. Rao, Molecular and biotechnological aspects of
xylanases, FEMS Microbiol. Rev. 23 (1999) 411456.
[20] S. Ahmed, S. Riaz, A. Jamil, Molecular cloning of fungal xylanases: an
overview, Appl. Microbiol. Biotechnol. 84 (2009) 1935.
[21] A. Knob, C. Terrasan, E. Carmona, -Xylosidases from lamentous fungi: an
overview, World J. Microbiol. Biotechnol. 26 (2010) 389407.
[22] T. Collins, C. Gerday, G. Feller, Xylanases, xylanase families and
extremophilic xylanases, FEMS Microbiol. Rev. 29 (2005) 323.
[23] D. Verma, T. Satyanarayana, Molecular approaches for ameliorating
microbial xylanases, Bioresour. Technol. 117 (2012) 360367.
[24] Z. Taibi, B. Saoudi, M. Boudelaa, H. Trigui, H. Belghith, A. Gargouri, A.
Ladjama, Purication and biochemical characterization of a highly
thermostable xylanase from Actinomadura sp. strain Cpt20 isolated from
poultry compost, Appl. Biochem. Biotechnol. 166 (2012) 663679.
[25] J.T.M. Buchert, A. Kantelinen, L. Viikari, Application of xylanases in the pulp
and paper industry, Bioresour. Technol. 50 (1995) 6572.
[26] V. Juturu, J.C. Wu, Microbial xylanases: engineering, production and
industrial applications, Biotechnol. Adv. 30 (2012) 12191227.
[27] F.J. St John, D.K. Godwin, J.F. Preston, E. Pozharski, J.C. Hurlbert,
Crystallization and crystallographic analysis of Bacillus subtilis xylanase C,
Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 65 (2009) 499503.
[28] D. De Vos, T. Collins, W. Nerinckx, S.N. Savvides, M. Claeyssens, C. Gerday, G.
Feller, J. Van Beeumen, Oligosaccharide binding in family 8 glycosidases:
crystal structures of active-site mutants of the beta-1, 4-xylanase pXyl from
Pseudoaltermonas haloplanktis TAH3a in complex with substrate and
product, Biochemistry 45 (2006) 47974807.
[29] C.R. Santos, A.N. Meza, Z.B. Hoffmam, J.C. Silva, T.M. Alvarez, R. Ruller, G.M.
Giesel, H. Verli, F.M. Squina, R.A. Prade, M.T. Murakami, Thermal-induced
[32]
[33]
[34]
[35]
[36]
[37]
[38]
[39]
[40]
[41]
[42]
[43]
[44]
[45]
[46]
[47]
[48]
[49]
[50]
[51]
[52]
[53]
conformational changes in the product release area drive the enzymatic
activity of xylanases 10B: crystal structure, conformational stability and
functional characterization of the xylanase 10B from Thermotoga petrophila
RKU-1, Biochem. Biophys. Res. Commun. 403 (2010) 214219.
M.D. Joshi, G. Sidhu, I. Pot, G.D. Brayer, S.G. Withers, L.P. McIntosh, Hydrogen
bonding and catalysis: a novel explanation for how a single amino acid
substitution can change the pH optimum of a glycosidase, J. Mol. Biol. 299
(2000) 255279.
Q. Han, N. Liu, H. Robinson, L. Cao, C. Qian, Q. Wang, L. Xie, H. Ding, Y. Huang,
J. Li, Z. Zhou, Biochemical characterization and crystal structure of a GH10
xylanase from termite gut bacteria reveal a novel structural feature and
signicance of its bacterial Ig-like domain, Biotechnol. Bioeng. 110 (2013)
30933103.
M.A. Sainz-Polo, S.V. Valenzuela, B. Gonzalez, F.I. Pastor, J. Sanz-Aparicio,
Structural analysis of glucuronoxylan-specic Xyn30D and its attached
CBM35 domain gives insights into the role of modularity in specicity, J.
Biol. Chem. 289 (2014) 3108831101.
S. Mohana, A. Shah, J. Divecha, D. Madamwar, Xylanase production by
Burkholderia sp. DMAX strain under solid state fermentation using distillery
spent wash, Bioresour. Technol. 99 (2008) 75537564.
B. Battan, J. Sharma, S.S. Dhiman, R.C. Kuhad, Enhanced production of
cellulasefree thermostable xylanase by Bacillus pumilus ASH and its
potential application in paper industry, Enzyme Microb. Technol. 41 (2008)
733739.
C.A. Poorna, P. Prema, Production of cellulase-free endoxylanase from novel
alkalophilic thermotolerant Bacillus pumilus by solid-state fermentation and
its application in wastepaper recycling, Bioresour. Technol. 98 (2007)
485490.
S.L. Wang, Y.H. Yen, I.L. Shih, A.C. Chang, W.T. Chang, W.C. Wu, Y.D. Chai,
Production of xylanases from rice bran by Streptomyces actuosus A-151,
Enzyme Microb. Technol. 33 (2003) 917925.
S. Ninawe, M. Kapoor, R.C. Kuhad, Purication and characterization of
extracellular xylanase from Streptomyces cyaneus SN32, Bioresour. Technol.
99 (2008) 12521258.
A. Ruiz-Arribas, J.M. Fernndez-Abalos, P. Snchez, A. Garda, R.I. Santamara,
Overproduction, purication, and biochemical characterization of a
xylanase (Xys1) from Streptomyces halstedii JM8, Appl. Environ. Microbiol. 6
(1995) 24142419.
Q.J. Yan, S.S. Hao, Z.Q. Jiang, Q. Zhai, W.W. Chen, Properties of a xylanase
from Streptomyces matensis being suitable for xylooligosaccharides
production, J. Mol. Catal. B: Enzym. 58 (2009) 7277.
S. Kaneko, A. Kuno, M. Muramatsu, S. Iwamatsu, I. Kusakabe, K. Hayashi,
Purication and characterization of a family G/11-xylanase from
Streptomyces olivaceoviridis E-86, Biosci. Biotechnol. Biochem. 64 (2000)
447451.
Y.R. Wang, H.L. Zhang, H.Z. He, H.Y. Luo, B. Yao, Characterization, gene
cloning and expression of a novel xylanase XYNB from Streptomyces
olivaceoviridis A1, Aquaculture 267 (2007) 328334.
C.G. Anthony, W.J. Thomas, Production, purication, and characterization of
beta (1,4)-endoxylanase of Streptomyces roseiscleroticus, Appl. Environ.
Microbiol. 57 (1991) 987992.
H. Tsujibo, K. Miyamoto, T. Kuda, K. Minami, T. Sakamoto, T. Hasegawa, Y.
Inamori, Purication, properties and partial amino acid sequences of
thermostable xylanase from Streptomyces thermoviolaceus OPC-520, Appl.
Microbiol. Biotechnol. 58 (1992) 371375.
T.S. Magnuson, D.L. Crawford, Purication and characterization of an
alkaline xylanase from Streptomyces viridosporus T7A, Enzyme Microb.
Technol. 21 (1997) 160164.
G.T. Dobreva, I.G. Pishtiyski, V.S. Stanchev, R. Mircheva, Optimization of
nutrient medium containing agricultural wastes for xylanase production by
Aspergillus niger B03 using optimal composite experimental design,
Bioresour. Technol. 28 (2007) 26712678.
Q.J. Yan, L. Wang, Z.Q. Jiang, S.Q. Yang, H.F. Zhu, L.T. Li, A xylosetolerant
beta-xylosidase from Paecilomyces thermophila: characterization and its
co-action with the endogenous xylanase, Bioresour. Technol. 99 (2008)
54025410.
M. Azin, R. Moravej, D. Zareh, Production of xylanase by Trichoderma
longibrachiatum on a mixture of wheat bran and wheat straw: optimization
of culture condition by Taguchi method, Enzyme Microb. Technol. 40 (2007)
801805.
X.T. Li, Z.Q. Jiang, L.T. Li, S.Q. Yang, W.Y. Feng, J.Y. Fan, I. Kusakabe,
Characterization of a cellulase-free, neutral xylanase from Thermomyces
lanuginosus CBS 288.54 and its biobleaching effect on wheat straw pulp,
Bioresour. Technol. 96 (2005) 13701379.
G.W. Harris, R.W. Pickersgill, I. Connerton, P. Debeire, J.P. Touzel, C. Breton,
S. Perez, Structural basis of the properties of an industrially relevant
thermophilic xylanase, Proteins 29 (1997) 7786.
I. Lasa, J. Berenguer, Thermophilic enzymes and their biotechnological
potential, Microbiology 9 (1993) 7789.
M. Ishihara, S. Tawata, S. Toyama, Purication and some properties of a
thermostable xylanase from thermophilic fungus strain HG-1, J. Ferment.
Bioeng. 83 (1997) 478480.
P.A. Dalby, Engineering enzymes for biocatalysis, Recent Pat. Biotechnol. 1
(2007) 19.
M.T. Reetz, J.D. Carballeira, Iterative saturation mutagenesis (ISM) for rapid
directed evolution of functional enzymes, Nat. Protoc. 2 (2007) 891903.
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
[54] D. Driss, F. Bhiri, R. Ghorbel, S.E. Chaabouni, Cloning and constitutive
expression of His-tagged xylanase GH 11 from Penicillium occitanis Pol6 in
Pichia pastoris X33: purication and characterization, Protein Expr. Purif. 83
(2012) 814.
[55] L. Mao, P. Meng, C. Zhou, L. Ma, G. Zhang, Y. Ma, Molecular cloning and
heterologous expression of an acid stable xylanase gene from Alternaria sp.
HB186, World J. Microbiol. Biotechnol. 28 (2012) 777784.
[56] S. Luetz, L. Giver, J. Lalonde, Engineered enzymes for chemical production,
Biotechnol. Bioeng. 101 (2008) 647653.
[57] S. Sen, V. Venkata Dasu, B. Mandal, Developments in directed evolution for
improving enzyme functions, Appl. Biochem. Biotechnol. 143 (2007)
212223.
[58] L. Song, A. Tsang, M. Sylvestre, Engineering a thermostable fungal GH10
xylanase, importance of N-terminal amino acids, Biotechnol. Bioeng. 112
(2015) 10811091.
[59] J.A. Diogo, Z.B. Hoffmam, L.M. Zanphorlin, J. Cota, C.B. Machado, L.D. Wolf, F.
Squina, A.R. Damasio, M.T. Murakami, R. Ruller, Development of a chimeric
hemicellulase to enhance the xylose production and thermotolerance,
Enzyme Microb. Technol. 69 (2015) 3137.
[60] H. Zhang, J. Li, J. Wang, Y. Yang, M. Wu, Determinants for the improved
thermostability of a mesophilic family 11 xylanase predicted by
computational methods, Biotechnol. Biofuels 7 (2014) 3.
[61] N.P. McHunu, S. Singh, K. Permaul, Expression of an alkalo-tolerant fungal
xylanase enhanced by directed evolution in Pichia pastoris and
Escherichia coli, J. Biotechnol. 141 (2009) 2630.
[62] M.L. Ludwiczek, I. DAngelo, G.N. Yalloway, J.A. Brockerman, M. Okon, J.E.
Nielsen, N.C. Strynadka, S.G. Withers, L.P. McIntosh, Strategies for
modulating the pH-dependent activity of a family 11 glycoside hydrolase,
Biochemistry 52 (2013) 31383156.
[63] C.A. Hokanson, G. Cappuccilli, T. Odineca, M. Bozic, C.A. Behnke, M. Mendez,
W.J. Coleman, R. Crea, Engineering highly thermostable xylanase variants
using an enhanced combinatorial library method, Protein Eng. Des. Sel. 24
(2011) 597605.
[64] P. Biely, Biochemical aspects of the production of microbial hemicellulases,
in: Hemicellulose and Hemicellulases, 1993, pp. 2951.
[65] A.C. Colabardini, A.C. Humanes, P.F. Gouvea, M. Savoldi, M.H.S. Goldman,
M.R.V.Z. Kress, . Bayram, J.V.D.C. Oliveira, M.D. Gomes, G.H. Braus, G.H.
Goldman, Molecular characterization of the Aspergillus nidulans fbxA
encoding an F-box protein involved in xylanase induction, Fungal Genet.
Biol. 49 (2012) 130140.
[66] A.R. Stricker, R.L. Mach, L.H. de Graaff, Regulation of transcription of
cellulases- and hemicellulases-encoding genes in Aspergillus niger and
Hypocrea jecorina (Trichoderma reesei), Appl. Microbiol. Biotechnol. 78
(2008) 211220.
[67] L.H. Degraaff, H.C. van den Broeck, A.J. van Ooijen, J. Visser, Regulation of the
xylanase-encoding xlnA gene of Aspergillus tubigensis, Mol. Microbiol. 12
(1994) 479490.
[68] G.J. Ruijter, J. Visser, Carbon repression in Aspergilli, FEMS Microbiol. Lett.
151 (1997) 103114.
[69] P. Kulmburg, M. Mathieu, C. Dowzer, J. Kelly, B. Felenbok, Specic binding
sites in the alcR and alcA promoters of the ethanol regulon for the CREA
repressor mediating carbon catabolite repression in Aspergillus nidulans,
Mol. Microbiol. 7 (1993) 847857.
[70] B. Cubero, C. Scazzocchio, Two different, adjacent and divergent zinc nger
binding sites are necessary for CREA-mediated carbon catabolite repression
in the proline gene cluster of Aspergillus nidulans, EMBO J. 13 (1994)
407415.
[71] J. Strauss, R.L. Mach, S. Zeilinger, G. Hartler, G. Stofer, M. Wolschek, C.P.
Kubicek, Cre1, the carbon catabolite repressor protein from Trichoderma
reesei, FEBS Lett. 376 (1995) 103107.
[72] A. Hershko, A. Ciechanover, The ubiquitin system, Annu. Rev. Biochem. 67
(1998) 425479.
[73] D. Skowyra, K.L. Craig, M. Tyers, S.J. Elledge, J.W. Harper, F-box proteins are
receptors that recruit phosphorylated substrates to the SCF ubiquitin-ligase
complex, Cell 91 (1997) 209219.
[74] W. Jonkers, M. Rep, Lessons from fungal F-box proteins, Eukaryot. Cell 8
(2009) 677695.
[75] S. Krappmann, N. Jung, B. Medic, S. Busch, R.A. Prade, G.H. Braus, The
Aspergillus nidulans F-box protein GrrA links SCF activity to meiosis, Mol.
Microbiol. 61 (2006) 7688.
[76] B. Bhunia, D. Dutta, S. Chaudhuri, Selection of suitable carbon, nitrogen and
sulphate source for the production of alkaline protease by Bacillus
licheniformis NCIM-2042, Not. Sci. Biol. 2 (2010) 5659.
[77] V. Betina, Bioactive Secondary Metabolite of Microorganisms. Process in
Industrial Microbiology, Elsevier Science, Amsterdam & New York, 1994.
[78] F.G. Priest, Extracellular enzyme synthesis in the genus Bacillus, Bacteriol.
Rev. 41 (1977) 711753.
[79] C.G. Kumar, R.K. Malik, M.P. Tiwari, K.D. Jany, Optimal production of Bacillus
alkaline protease using a cheese whey medium, Microbiol. Aliment. Nutr. 17
(1999) 3948.
[80] O. Garcia-Kirchner, M. Munoz-Aguilar, R. Perez-Villalva, C. Huitron-Vargas,
Mixed submerged fermentation with two lamentous fungi for cellulolytic
and xylanolytic enzyme production, Appl. Biochem. Biotechnol. 98100
(2002) 11051114.
1053
[81] P. Tallapragada, K. Venkatesh, Isolation, identication and optimization of
xylanase enzyme produced by Aspergillus niger under submerged
fermentation, J. Microbiol. Biotechnol. Res. 1 (2011) 137147.
[82] S.H. Moon, S.J. Parulekar, A parametric study ot protease production in batch
and fed-batch cultures of Bacillus rmus, Biotechnol. Bioeng. 37 (1991)
467483.
[83] S.F.G. Oskouie, F. Tabandeh, B. Yakhchali, F. Eftekhar, Enhancement of
alkaline protease production by Bacillus clausii using Taguchi experimental
design, Afr. J. Biotechnol. 6 (2007) 25592564.
[84] M. Muller, F. Segato, R.A. Prade, M.R. Wilkins, High-yield recombinant
xylanase production by Aspergillus nidulans under pyridoxine limitation, J.
Ind. Microbiol. Biotechnol. 41 (2014) 15631570.
[85] W. Mao, R. Pan, D. Freedman, High production of alkaline protease by
Bacillus licheniformis in a fed-batch fermentation using a synthetic medium,
J. Ind. Microbiol. 11 (1992) 16.
[86] N. Fujiwara, K. Yamamoto, A. Masui, Utilization of a thermostable alkaline
protease from an alkalophilic thermophile for the recovery of silver from
used X-ray lm, J. Ferment. Bioeng. 72 (1991) 306308.
[87] U. Hbner, U. Bock, K. Schgerl, Production of alkaline serine protease
subtilisin Carlsberg by Bacillus licheniformis on complex medium in a stirred
tank reactor, Appl. Microbiol. Biotechnol. 40 (1993) 182188.
[88] R. Wanga, R.C.S. Lawb, C. Webba, Protease production and conidiation by
Aspergillus oryzae in our fermentation, Process Biochem. 40 (2005)
217227.
[89] M. Kapoor, L.M. Nair, R.C. Kuhad, Cost-effective xylanase production from
free and immobilized Bacillus pumilus strain MK001 and its application in
saccharication of Prosopis juliora, Biochem. Eng. J. 38 (2008) 8897.
[90] U. Holker, A. Jurgen, Solid-state fermentation e are there any
biotechnological advantages, Curr. Opin. Microbiol. 8 (2005) 301306.
[91] A. Pandey, P. Selvakumar, C.R. Soccol, P. Nigam, Solid State fermentation for
the production of industrial enzymes, Curr. Sci. 77 (1999) 149162.
[92] N.C. de Alencar Guimaraes, M. Sorgatto, C. Peixoto-Nogueira Sde, J.H. Betini,
F.F. Zanoelo, M.R. Marques, L. de Moraes Polizeli Mde, G.C. Giannesi,
Bioprocess and biotecnology: effect of xylanase from Aspergillus niger and
Aspergillus avus on pulp biobleaching and enzyme production using
agroindustrial residues as substract, Springerplus 2 (2013) 380.
[93] S.K. Ang, E.M. Shaza, Y. Adibah, A.A. Suraini, M.S. Madihah, Production of
cellulases and xylanase by Aspergillus fumigatus SK1 using untreated oil
palm trunk through solid state fermentation, Process Biochem. 48 (2013)
12931302.
[94] A. Sadaf, S.K. Khare, Production of Sporotrichum thermophile xylanase by
solid state fermentation utilizing deoiled Jatropha curcas seed cake and its
application in xylooligosachharide synthesis, Bioresour. Technol. 153 (2014)
126130.
[95] B. Biswanath, P. Bhattacharya, Kinetic studies of alkaline protease from
Bacillus licheniformis NCIM-2042, J. Microbiol. Biotechnol. 22 (2012)
17581766.
[96] B. Bhunia, A. Dey, Statistical approach for optimization of physiochemical
requirements on alkaline protease production from Bacillus licheniformis
NCIM 2042, Enzyme Res. 2012 (2012).
[97] D. Verma, T. Satyanarayana, Production of cellulase-free xylanase by the
recombinant Bacillus subtilis and its applicability in paper pulp bleaching,
Biotechnol. Prog. 29 (2013) 14411447.
[98] G.M. Maciel, L.P.D.S. Vandenberghe, C.W.I. Haminiuk, R.C. Fendrich, B.E.D.
Bianca, T.Q.D.S. Brandalize, A. Pandey, C.R. Soccol, Xylanase production by
Aspergillus niger LPB 326 in solid-state fermentation using statistical
experimental designs, Food Technol. Biotechnol. 46 (2008) 183189.
[99] P. Calk, G. Calk, T.H. zdamar, Bioprocess development for serine alkaline
protease production: a review, Rev. Chem. Eng. 17 (2001) 162.
[100] B. Bhunia, B. Basak, P. Bhattacharya, A. Dey, Process engineering studies to
investigate the effect of temperature and pH on kinetic parameters of
alkaline protease production, J. Biosci. Bioeng. 115 (2013) 8689.
[101] R.A. Abusham, R.N. Rahman, A.B. Salleh, M. Basri, Optimization of physical
factors affecting the production of thermo-stable organic solvent-tolerant
protease from a newly isolated halo tolerant Bacillus subtilis strain Rand,
Microb. Cell Fact. 8 (2009) 20.
[102] C.T. Calam, Process Development in Antibiotic Fermentations, Cambridge
University Press, Cambridge, 1987.
[103] B. Kristiansen, M. Matteey, J. Linden, Citric Acid Biotechnology, Taylor and
Francis, London, 1999.
[104] M. Papagianni, M. Moo-Young, Protease secretion in glucoamylase producer
Aspergillus niger cultures: fungal morphology and inoculum effects, Process
Biochem. 37 (2002) 12711278.
[105] L.F. Dobson, C.C. OCleirigh, D.G. OShea, The inuence of morphology on
geldanamycin production in submerged fermentations of Streptomyces
hygroscopicus var. geldanus, Appl. Microbiol. Biotechnol. 79 (2008) 859866.
[106] J.C. van Suijdam, N.W.F. Kossen, P.G. Paul, An inoculum technique for the
production of fungal pellets, Eur. J. Appl. Microbiol. Biotechnol. 10 (1980)
211221.
[107] V. Gancheva, T. Dimova, [Biosynthesis of gibberellins. II. Effect of the amount
and age of the inoculum on gibberellin biosynthesis by the strain Fusarium
moniliforme IM-11], Acta Microbiol. Bulg. 14 (1984) 7479.
[108] S.E. Vecht-Lifshitz, S. Magdassi, S. Braun, Pellet formation and cellular
aggregation in Streptomyces tendae, Biotechnol. Bioeng. 35 (1990) 890896.
1054
U.S.P. Uday et al. / International Journal of Biological Macromolecules 82 (2016) 10411054
[109] V.S. Kranthi, D.M. Rao, P. Jaganmohan, Production of protease by Aspergillus
avus through solid state fermentation using different oil seed cakes, Int. J.
Microbiol. Res. 3 (2012) 1215.
[110] J. Nielsen, J. Villadsen, Bioreaction Engineering Principles, Plenium, New
York, 1994.
[111] J. Chaloupka, Temperature as a factor regulating the synthesis of microbial
enzymes, Microbiol. Sci. 2 (1985) 8690.
[112] J. Frankena, G.M. Koningstein, H.W. van Verseveld, A.H. Stouthamer, Effect
of different limitations in chemostat cultures on growth and production of
exocellular protease by Bacillus licheniformis, Appl. Microbiol. Biotechnol.
(1986) 106112.
[113] M. Michelin, L. Polizeli Mde, D.P. Silva, D.S. Ruzene, A.A. Vicente, J.A. Jorge,
H.F. Terenzi, J.A. Teixeira, Production of xylanolytic enzymes by Aspergillus
terricola in stirred tank and airlift tower loop bioreactors, J. Ind. Microbiol.
Biotechnol. 38 (2011) 19791984.
[114] K. Schugerl, S.R. Gerlach, D. Siedenberg, Inuence of the process parameters
on the morphology and enzyme production of Aspergilli, Adv. Biochem. Eng.
Biotechnol. 60 (1998) 195266.
[115] I. Michalik, E. Szabova, A. Polakova, D. Urminska, The selection of Bacillus
licheniformis strains for protease production: characterization of bacterial
alkaline protease, Biologia (Bratisl.) 50 (1995) 249252.
[116] A. Hille, T.R. Neu, D.C. Hempel, H. Horn, Oxygen proles and biomass
distribution in biopellets of Aspergillus niger, Biotechnol. Bioeng. 92 (2005)
614623.
[117] R. Biswas, V. Sahai, S. Mishra, V.S. Bisaria, Bioprocess strategies for enhanced
production of xylanase by Melanocarpus albomyces IITD3A on agro-residual
extract, J. Biosci. Bioeng. 110 (2010) 702708.
[118] P.J. Punt, A. Levasseur, H. Visser, J. Wery, E. Record, Fungal protein
production: design and production of chimeric proteins, Annu. Rev.
Microbiol. 65 (2011) 5769.
[119] A. Ganguly, P.K. Chatterjee, A. Dey, Studies on ethanol production from
water hyacinth A review, Renew. Sustain. Energy Rev. 16 (2012) 966972.
[120] A.L. Banka, S.A. Guralp, E. Gulari, Secretory expression and characterization
of two hemicellulases, xylanase, and beta-xylosidase, isolated from Bacillus
subtilis M015, Appl. Biochem. Biotechnol. 174 (2014) 27022710.
[121] G. Banerjee, S. Car, J.S. Scott-Craig, M.S. Borrusch, J.D. Walton, Rapid
optimization of enzyme mixtures for deconstruction of diverse
pretreatment/biomass feedstock combinations, Biotechnol. Biofuels 3
(2010) 22.
[122] J.B. Kristensen, C. Felby, H. Jorgensen, Yield-determining factors in
high-solids enzymatic hydrolysis of lignocellulose, Biotechnol. Biofuels 2
(2009) 11.
[123] M. Chen, J. Zhao, L. Xia, Enzymatic hydrolysis of maize straw polysaccharides
for the production of reducing sugars, Carbohydr. Polym. 71 (2008)
411415.
[124] J.B. Kristensen, C. Felby, H. Jorgensen, Determining yields in high solids
enzymatic hydrolysis of biomass, Appl. Biochem. Biotechnol. 156 (2009)
557562.
[125] G. Banerjee, J.S. Scott-Craig, J.D. Walton, Improving enzymes for biomass
conversion: a basic research perspective, BioEnergy Res. 3 (2010) 8292.
[126] Q. Qing, C.E. Wyman, Hydrolysis of different chain length xylooliogmers by
cellulase and hemicellulase, Bioresour. Technol. 102 (2011) 13591366.
[127] B. Sipos, Z. Benko, D. Dienes, K. Reczey, L. Viikari, M. Siika-aho,
Characterisation of specic activities and hydrolytic properties of
cell-wall-degrading enzymes produced by Trichoderma reesei Rut C30 on
different carbon sources, Appl. Biochem. Biotechnol. 161 (2010) 347364.
[128] Z.X. Lin, H.M. Zhang, X.J. Ji, J.W. Chen, H. Huang, Hydrolytic enzyme of
cellulose for complex formulation applied research, Appl. Biochem.
Biotechnol. 164 (2011) 2333.
[129] H. Ravalason, I. Herpol-Gimbert, E. Record, F. Bertaud, S. Grisel, Fusion of a
family 1 carbohydrate binding module to the Pycnoporus cinnabarinus
laccase for efcient softwood kraft pulp biobleaching, J. Biotechnol. 142
(2009) 220226.
[130] T.A. Pham, J.G. Berrin, E. Record, A. To Kim, J.C. Sigoillot, Hydrolysis of
softwood by Aspergillus mannanase: role of a carbohydrate binding module,
J. Biotechnol. 148 (2010) 163170.
[131] R. Araki, M.K. Ali, M. Sakka, T. Kimura, K. Sakka, K. Ohmiya, Essential role of
the family-22 carbohydrate-binding modules for beta-1,3-1,4-glucanase
activity of Clostridium stercorarium Xyn10B, FEBS Lett. 561 (2004) 155158.
[132] R. Araki, S. Karita, A. Tanaka, T. Kimura, K. Sakka, Effect of family 22
carbohydrate-binding module on the thermostability of Xyn10B catalytic
module from Clostridium stercorarium, Biosci. Biotechnol. Biochem. 70
(2006) 30393041.
[133] I. Benoit, M. Asther, G. Sulzenbacher, E. Record, L. Marmuse, G. Parsiegla, I.
Gimbert, C. Bignon, Respective importance of protein folding and
glycosylation in the thermal stability of recombinant feruloyl esterase A,
FEBS Lett. 580 (2006) 58155821.
Vous aimerez peut-être aussi
- English English: Bring P5.00 Small Bag Only Bring P5.00 Small Bag OnlyDocument1 pageEnglish English: Bring P5.00 Small Bag Only Bring P5.00 Small Bag Onlya3kt95Pas encore d'évaluation
- Perfect AttendanceDocument13 pagesPerfect Attendancea3kt95Pas encore d'évaluation
- 2nd Qrtr. PointersDocument1 page2nd Qrtr. Pointersa3kt95Pas encore d'évaluation
- Adriane T. Taban Adriane T. TabanDocument3 pagesAdriane T. Taban Adriane T. Tabana3kt95Pas encore d'évaluation
- Count The Objects and Write The Correct Answer. (0, 1, 2, 3, 4, 5)Document1 pageCount The Objects and Write The Correct Answer. (0, 1, 2, 3, 4, 5)a3kt95Pas encore d'évaluation
- Mixing Tank Hammer Mill Jaw Crusher: Bamboo Poles 1 M Ammonium PersulfateDocument1 pageMixing Tank Hammer Mill Jaw Crusher: Bamboo Poles 1 M Ammonium Persulfatea3kt95Pas encore d'évaluation
- Count The Objects and Write The Correct Answer. (0, 1, 2, 3, 4, 5)Document1 pageCount The Objects and Write The Correct Answer. (0, 1, 2, 3, 4, 5)a3kt95Pas encore d'évaluation
- FPPS 183Document3 pagesFPPS 183a3kt95Pas encore d'évaluation
- First Sem SchedDocument1 pageFirst Sem Scheda3kt95Pas encore d'évaluation
- Application Form 2016: Personal InformationDocument3 pagesApplication Form 2016: Personal Informationa3kt95Pas encore d'évaluation
- Criteria For Qualification:: ST NDDocument8 pagesCriteria For Qualification:: ST NDa3kt95Pas encore d'évaluation
- CEAT+Thesis Practicum Manuscript Formatting Guidelines Revised+July+2013 PDFDocument87 pagesCEAT+Thesis Practicum Manuscript Formatting Guidelines Revised+July+2013 PDFa3kt95Pas encore d'évaluation
- ENSC 26 Course Outline - Lecture - Rev - 2016!17!1st - SemDocument2 pagesENSC 26 Course Outline - Lecture - Rev - 2016!17!1st - Sema3kt95Pas encore d'évaluation
- ENSC 26 Course Outline - Laboratory - Rev - 2015!16!1st - SemDocument2 pagesENSC 26 Course Outline - Laboratory - Rev - 2015!16!1st - Sema3kt95Pas encore d'évaluation
- ChE 154 Lecture 1Document35 pagesChE 154 Lecture 1a3kt95Pas encore d'évaluation
- PipingDocument1 pagePipinga3kt95Pas encore d'évaluation
- (Answer: V 30.5781 KN, M 112.8374 KNM) : Problem Set 4Document2 pages(Answer: V 30.5781 KN, M 112.8374 KNM) : Problem Set 4a3kt95Pas encore d'évaluation
- Problem Set 4 (ES 11 - I) : CM I CM y CM XDocument3 pagesProblem Set 4 (ES 11 - I) : CM I CM y CM Xa3kt95Pas encore d'évaluation
- And A Rocker at V.: Problem Set 2Document3 pagesAnd A Rocker at V.: Problem Set 2a3kt95Pas encore d'évaluation
- Shoe Dog: A Memoir by the Creator of NikeD'EverandShoe Dog: A Memoir by the Creator of NikeÉvaluation : 4.5 sur 5 étoiles4.5/5 (537)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeD'EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeÉvaluation : 4 sur 5 étoiles4/5 (5795)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceD'EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceÉvaluation : 4 sur 5 étoiles4/5 (895)
- The Yellow House: A Memoir (2019 National Book Award Winner)D'EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Évaluation : 4 sur 5 étoiles4/5 (98)
- Grit: The Power of Passion and PerseveranceD'EverandGrit: The Power of Passion and PerseveranceÉvaluation : 4 sur 5 étoiles4/5 (588)
- The Little Book of Hygge: Danish Secrets to Happy LivingD'EverandThe Little Book of Hygge: Danish Secrets to Happy LivingÉvaluation : 3.5 sur 5 étoiles3.5/5 (400)
- The Emperor of All Maladies: A Biography of CancerD'EverandThe Emperor of All Maladies: A Biography of CancerÉvaluation : 4.5 sur 5 étoiles4.5/5 (271)
- Never Split the Difference: Negotiating As If Your Life Depended On ItD'EverandNever Split the Difference: Negotiating As If Your Life Depended On ItÉvaluation : 4.5 sur 5 étoiles4.5/5 (838)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyD'EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyÉvaluation : 3.5 sur 5 étoiles3.5/5 (2259)
- On Fire: The (Burning) Case for a Green New DealD'EverandOn Fire: The (Burning) Case for a Green New DealÉvaluation : 4 sur 5 étoiles4/5 (74)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureD'EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureÉvaluation : 4.5 sur 5 étoiles4.5/5 (474)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryD'EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryÉvaluation : 3.5 sur 5 étoiles3.5/5 (231)
- Team of Rivals: The Political Genius of Abraham LincolnD'EverandTeam of Rivals: The Political Genius of Abraham LincolnÉvaluation : 4.5 sur 5 étoiles4.5/5 (234)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaD'EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaÉvaluation : 4.5 sur 5 étoiles4.5/5 (266)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersD'EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersÉvaluation : 4.5 sur 5 étoiles4.5/5 (345)
- The Unwinding: An Inner History of the New AmericaD'EverandThe Unwinding: An Inner History of the New AmericaÉvaluation : 4 sur 5 étoiles4/5 (45)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreD'EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreÉvaluation : 4 sur 5 étoiles4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)D'EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Évaluation : 4.5 sur 5 étoiles4.5/5 (121)
- Her Body and Other Parties: StoriesD'EverandHer Body and Other Parties: StoriesÉvaluation : 4 sur 5 étoiles4/5 (821)
- Research Paper On Recombinant Dna TechnologyDocument7 pagesResearch Paper On Recombinant Dna Technologyvagipelez1z2100% (1)
- BISC403 Sample Exam 4 W - Answers 20SpDocument5 pagesBISC403 Sample Exam 4 W - Answers 20SpGrace MillsPas encore d'évaluation
- Biotechnology 101Document273 pagesBiotechnology 101Mitasha Bharadwaj AroraPas encore d'évaluation
- Class XII: Biology Chapter 6: Molecular Basis of InheritanceDocument9 pagesClass XII: Biology Chapter 6: Molecular Basis of Inheritancevishlesh parmarPas encore d'évaluation
- Animal Behavior GoodenoughDocument546 pagesAnimal Behavior GoodenoughDEBALIKA ROYPas encore d'évaluation
- Lecture Notes (Genetic Counselling)Document25 pagesLecture Notes (Genetic Counselling)Zearo GamingPas encore d'évaluation
- Agricultural MicrobiologyDocument3 pagesAgricultural MicrobiologyShahriar ShamimPas encore d'évaluation
- Caraga Regional Science High School: What I KnowDocument13 pagesCaraga Regional Science High School: What I KnowFaust HacklPas encore d'évaluation
- PCB3103C Exam 2 (Take-Home)Document5 pagesPCB3103C Exam 2 (Take-Home)Pmorris1898Pas encore d'évaluation
- Sex Linkage and Sex DeterminationDocument8 pagesSex Linkage and Sex DeterminationBrian MachachaPas encore d'évaluation
- Williamthomascvjan 2024Document3 pagesWilliamthomascvjan 2024api-438172507Pas encore d'évaluation
- The Origin of Viruses: Patrick Forterre, David PrangishviliDocument7 pagesThe Origin of Viruses: Patrick Forterre, David PrangishviliDumisani NguniPas encore d'évaluation
- MS397 Final MaalaDocument4 pagesMS397 Final MaalaGabie MaalaPas encore d'évaluation
- Promoter Nucleotide Transcribed RNA Polymerase: Operator - A Segment ofDocument3 pagesPromoter Nucleotide Transcribed RNA Polymerase: Operator - A Segment ofJitesh SoniPas encore d'évaluation
- Sample Question Paper Class XII (2019-20) BiologyDocument14 pagesSample Question Paper Class XII (2019-20) BiologyArif BhaiPas encore d'évaluation
- Q&A Modern Blood Banking Transfusion Practices - Harmening, Denise M.Document55 pagesQ&A Modern Blood Banking Transfusion Practices - Harmening, Denise M.Evan GaufoPas encore d'évaluation
- BCMB20002 MST2 Sem1 2016Document6 pagesBCMB20002 MST2 Sem1 2016kfqhnmh87hPas encore d'évaluation
- Chapter 11 - Gene ExpressionDocument2 pagesChapter 11 - Gene Expressionapi-272677238Pas encore d'évaluation
- Positional Cloning of The Wheat VernalizationDocument6 pagesPositional Cloning of The Wheat VernalizationvahidvasharPas encore d'évaluation
- 2005 Volume 96 Marine Biotechnology IDocument261 pages2005 Volume 96 Marine Biotechnology IabavoPas encore d'évaluation
- 3.2 The Principles and Mechanism of InheritanceDocument6 pages3.2 The Principles and Mechanism of InheritanceajakazPas encore d'évaluation
- Sample QuestionsDocument52 pagesSample QuestionsMiri HahiashvilliPas encore d'évaluation
- (2011) Production of Ant Proteins by Filamentous FungiDocument21 pages(2011) Production of Ant Proteins by Filamentous FungiAnthi KarnaouriPas encore d'évaluation
- Dna Sequencing ThesisDocument7 pagesDna Sequencing Thesisdwndnjfe100% (2)
- National Geographic UK - September 2019-CompressedDocument133 pagesNational Geographic UK - September 2019-CompressedEmily Tan67% (3)
- TOEFL Test 2Document23 pagesTOEFL Test 2Rio HandokoPas encore d'évaluation
- 2015 Molecular Biology of The Cell 6th Edition Test BankDocument36 pages2015 Molecular Biology of The Cell 6th Edition Test Bankgulessideslip.22kz100% (38)
- Molecular DiagnosticsDocument583 pagesMolecular DiagnosticsAnthony93% (14)
- Artículo. Taller 01Document8 pagesArtículo. Taller 01Karol Stephany Cifuentes SanchezPas encore d'évaluation
- Concept of GeneDocument30 pagesConcept of GeneSubhapam KunduPas encore d'évaluation