Académique Documents
Professionnel Documents
Culture Documents
Correspondence Analysis: Sample Statfolio: Sample Data
Transféré par
Anonymous FZNn6rBTitre original
Copyright
Formats disponibles
Partager ce document
Partager ou intégrer le document
Avez-vous trouvé ce document utile ?
Ce contenu est-il inapproprié ?
Signaler ce documentDroits d'auteur :
Formats disponibles
Correspondence Analysis: Sample Statfolio: Sample Data
Transféré par
Anonymous FZNn6rBDroits d'auteur :
Formats disponibles
STATGRAPHICS – Rev.
7/6/2009
Correspondence Analysis
The Correspondence Analysis procedure creates a map of the rows and columns in a two-way
contingency table for the purpose of providing insights into the relationships amongst the
categories of the row and/or column variables. Often, no more than two or three dimensions are
needed to display most of the variability or “inertia” in the table. An important part of the output
is a correspondence map on which the distance between two categories is a measure of their
similarity.
Sample StatFolio: correspondence.sgp
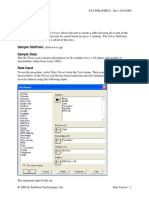
Sample Data:
The file funding.sgd contains data that describe the level of funding awarded to 796 scientific
researchers (from Greenacre, 2007). The researchers are divided into 10 scientific disciplines,
which form the rows of the table shown below:
A B C D E
Geology 3 19 39 14 19
Biochemistry 1 2 13 1 12
Chemistry 6 25 49 21 29
Zoology 3 15 41 35 26
Physics 10 22 47 9 26
Engineering 3 11 25 15 34
Microbiology 1 6 14 5 11
Botany 0 12 34 17 23
Statistics 2 5 11 4 7
Mathematics 2 11 37 8 20
The columns identify the level of funding awarded to each researcher, from A (most highly
funded) to D (least funded), with E representing no funding at all. In addition to the rows and
columns that will be used to generate the map, additional supplementary rows and columns may
be added to the table for plotting purposes only. For example, the table below adds two
supplementary rows and one supplementary column:
A B C D E Y
Geology 3 19 39 14 19 0
Biochemistry 1 2 13 1 12 1
Chemistry 6 25 49 21 29 0
Zoology 3 15 41 35 26 0
Physics 10 22 47 9 26 1
Engineering 3 11 25 15 34 1
Microbiology 1 6 14 5 11 1
Botany 0 12 34 17 23 1
Statistics 2 5 11 4 7 0
Mathematics 2 11 37 8 20 2
Museums 4 12 11 9 7
Math Sciences 4 16 48 12 27
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 1
STATGRAPHICS – Rev. 7/6/2009
Data Input
The procedure is designed to handle two types of data:
1. Data that has been tabulated into a two-way table such as that shown above.
2. Data that has not yet been tabulated.
For example, the data file or the above example might contain 792 rows, one for each researcher:
In that case, select the Untabulated (Observations) radio button on the data input dialog box and
specify the columns of the data file as shown below:
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 2
STATGRAPHICS – Rev. 7/6/2009
Row variable: name of numeric or non-numeric column identifying the row category for
each individual.
Column variable: name of numeric or non-numeric column identifying the column category
for each individual.
Select: subset selection.
On the other hand, the data file might contain tabulated counts and be laid out in a format similar
to the two-way table:
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 3
STATGRAPHICS – Rev. 7/6/2009
In that case, select the Tabulated (Counts) radio button and specify the names of the columns as
shown below:
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 4
STATGRAPHICS – Rev. 7/6/2009
Counts: two or more numeric columns containing the counts in the table.
Labels: column containing identifiers for each row of the table.
Select: subset selection.
Analysis Options
The Analysis Options dialog box is shown below:
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 5
STATGRAPHICS – Rev. 7/6/2009
Number of Dimensions to Extract: the number of dimensions that you wish to extract
from the table. You must specify a number between 2 and one less than the smaller of the
number of rows and the number of columns. Often, 2 or 3 dimensions is sufficient to
explain most of the differences amongst the categories.
Supplementary Points: the number of rows and/or columns that you wish to exclude
from the calculations. These rows and columns will be plotted on the correspondence
map but will not be used to determine the scaling.
Analysis Summary
The Analysis Summary displays the data in the form of a contingency table:
Correspondence Analysis
Contingency Table
A B C D E TOTAL
Geology 3 19 39 14 10 85
Biochemistry 1 2 13 1 12 29
Chemistry 6 25 49 21 29 130
Zoology 3 15 41 35 26 120
Physics 10 22 47 9 26 114
Engineering 3 11 25 15 34 88
Microbiology 1 6 14 5 11 37
Botany 0 12 34 17 23 86
Statistics 2 5 11 4 7 29
Mathematics 2 11 37 8 20 78
TOTAL 31 128 310 129 198 796
Supplemental Rows
A B C D E
Museums 4 12 11 19 7
Math Sciences 4 16 48 12 27
Supplemental Columns
Y
Geology 0
Biochemistry 1
Chemistry 0
Zoology 0
Physics 1
Engineering 1
Microbiology 1
Botany 1
Statistics 0
Mathematics 1
Any supplemental rows or columns will be displayed separately from the counts used to
calculate the scaling.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 6
STATGRAPHICS – Rev. 7/6/2009
Row and Column Profiles
An important concept in correspondence analysis is that of a row or column profile. A profile is
simply a row or column of a contingency table, divided by its total. The row and column profiles
for the sample data are shown below:
Row and Column Profiles
Row Profiles
A B C D E MASS
Geology 0.035 0.224 0.459 0.165 0.118 0.107
Biochemistry 0.034 0.069 0.448 0.034 0.414 0.036
Chemistry 0.046 0.192 0.377 0.162 0.223 0.163
Zoology 0.025 0.125 0.342 0.292 0.217 0.151
Physics 0.088 0.193 0.412 0.079 0.228 0.143
Engineering 0.034 0.125 0.284 0.170 0.386 0.111
Microbiology 0.027 0.162 0.378 0.135 0.297 0.046
Botany 0.000 0.140 0.395 0.198 0.267 0.108
Statistics 0.069 0.172 0.379 0.138 0.241 0.036
Mathematics 0.026 0.141 0.474 0.103 0.256 0.098
MASS 0.039 0.161 0.389 0.162 0.249
Column Profiles
A B C D E MASS
Geology 0.097 0.148 0.126 0.109 0.051 0.107
Biochemistry 0.032 0.016 0.042 0.008 0.061 0.036
Chemistry 0.194 0.195 0.158 0.163 0.146 0.163
Zoology 0.097 0.117 0.132 0.271 0.131 0.151
Physics 0.323 0.172 0.152 0.070 0.131 0.143
Engineering 0.097 0.086 0.081 0.116 0.172 0.111
Microbiology 0.032 0.047 0.045 0.039 0.056 0.046
Botany 0.000 0.094 0.110 0.132 0.116 0.108
Statistics 0.065 0.039 0.035 0.031 0.035 0.036
Mathematics 0.065 0.086 0.119 0.062 0.101 0.098
MASS 0.039 0.161 0.389 0.162 0.249
For example, the top left cell (Geology – A) represents approximately 3.5% of the Geology row
and approximately 9.7% of the A column.
The table also displays the total mass associated with each row and column. Mass is a measure of
the total count in a row or column divided by the total count in the entire table. For example,
about 10.7% of the researchers in the table are in Geology.
Pane Options
Specify the number of decimal places for displaying the profiles.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 7
STATGRAPHICS – Rev. 7/6/2009
Mosaic Plot
An interesting way to plot the data in a contingency table is through a Mosaic Plot:
Mosaic Plot
Geology
Biochemistry A
B
Chemistry
C
Zoology D
E
Physics
Engineering
Microbiology
Botany
Statistics
Mathematics
In this plot, the height of each row is proportional to its row total. The width of the bars within
each row is proportional to the frequency of each cell within that row (its row profile). This
results in bars whose area is proportional to the frequency in a particular cell.
Pane Options
Direction: the orientation of the bars. Switching to a vertical plot will plot column profiles
rather than row profiles.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 8
STATGRAPHICS – Rev. 7/6/2009
Expected Frequencies and Residuals
If the row and column classifications in a contingency table are independent, the expected count
in a cell is the total count in the table, multiplied by the row mass, multiplied by the column
mass. The Expected Frequencies and Residuals pane displays these expected values, together
with the differences between the actual counts and those expectations:
Expected Frequencies and Residuals
Expected Frequencies
A B C D E
Geology 3.31 13.67 33.10 13.78 21.14
Biochemistry 1.13 4.66 11.29 4.70 7.21
Chemistry 5.06 20.90 50.63 21.07 32.34
Zoology 4.67 19.30 46.73 19.45 29.85
Physics 4.44 18.33 44.40 18.47 28.36
Engineering 3.43 14.15 34.27 14.26 21.89
Microbiology 1.44 5.95 14.41 6.00 9.20
Botany 3.35 13.83 33.49 13.94 21.39
Statistics 1.13 4.66 11.29 4.70 7.21
Mathematics 3.04 12.54 30.38 12.64 19.40
Residuals (Observed - Expected)
A B C D E
Geology -0.31 5.33 5.90 0.22 -11.14
Biochemistry -0.13 -2.66 1.71 -3.70 4.79
Chemistry 0.94 4.10 -1.63 -0.07 -3.34
Zoology -1.67 -4.30 -5.73 15.55 -3.85
Physics 5.56 3.67 2.60 -9.47 -2.36
Engineering -0.43 -3.15 -9.27 0.74 12.11
Microbiology -0.44 0.05 -0.41 -1.00 1.80
Botany -3.35 -1.83 0.51 3.06 1.61
Statistics 0.87 0.34 -0.29 -0.70 -0.21
Mathematics -1.04 -1.54 6.62 -4.64 0.60
For example, the expected count in the upper left corner of the table if Discipline and Funding
are independent is 3.31. Subtracting 3.31 from the observed value of 3 leaves a residual equal to
-0.31.
Pane Options
Specify the number of decimal places for displaying the frequencies and residuals.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 9
STATGRAPHICS – Rev. 7/6/2009
Chi-Square Distances
A well-known statistic for measuring how much the counts in a contingency table differ from
that expected if the row and column classifications were independent is given by the chi-square
statistic. If eij represents the expected count in the cell for row i, column j and nij represents the
actual count, then the chi-square statistic is calculated from:
r c (nij eij ) 2
2 (1)
i 1 j 1 eij
The table below shows the contributions of each cell to the chi-square statistic:
Chi-Square Distances
A B C D E TOTAL
Geology 0.029 2.080 1.050 0.004 5.873 9.036
Biochemistry 0.015 1.521 0.258 2.913 3.176 7.882
Chemistry 0.173 0.802 0.052 0.000 0.344 1.373
Zoology 0.599 0.957 0.703 12.438 0.496 15.194
Physics 6.964 0.734 0.153 4.859 0.196 12.906
Engineering 0.053 0.702 2.508 0.038 6.700 10.001
Microbiology 0.135 0.000 0.012 0.166 0.351 0.663
Botany 3.349 0.242 0.008 0.673 0.121 4.393
Statistics 0.671 0.024 0.008 0.104 0.006 0.814
Mathematics 0.354 0.190 1.444 1.704 0.018 3.710
TOTAL 12.343 7.252 6.196 22.899 17.282 65.972
The cell with the largest contribution to the total chi-squared statistic is displayed in red. The
combined chi-squared contribution of all cells in a row or column (shown under TOTAL) can
also be thought of as the distance between an observed row or column profile and the centroid of
the all the row or column profiles, where the centroid is the profile expected under the
assumption of independence.
Relative Inertias
If the chi-square distances are divided by the total count in the contingency table, then the
resulting quantities are called inertia. If the inertia in a cell is further divided by the total inertia
in the entire table, the result is called a relative inertia. Summing over a row or column yields the
relative inertia of that row or column. The table below shows the relative inertias of each cell,
row and column in the contingency table for the sample data:
Relative Inertias
A B C D E TOTAL
Geology 0.000 0.032 0.016 0.000 0.089 0.137
Biochemistry 0.000 0.023 0.004 0.044 0.048 0.119
Chemistry 0.003 0.012 0.001 0.000 0.005 0.021
Zoology 0.009 0.015 0.011 0.189 0.008 0.230
Physics 0.106 0.011 0.002 0.074 0.003 0.196
Engineering 0.001 0.011 0.038 0.001 0.102 0.152
Microbiology 0.002 0.000 0.000 0.003 0.005 0.010
Botany 0.051 0.004 0.000 0.010 0.002 0.067
Statistics 0.010 0.000 0.000 0.002 0.000 0.012
Mathematics 0.005 0.003 0.022 0.026 0.000 0.056
TOTAL 0.187 0.110 0.094 0.347 0.262 1.000
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 10
STATGRAPHICS – Rev. 7/6/2009
Note that the cell shown in red contributes about 18.9% of the total inertia in the contingency
table.
Inertia and Chi-Square Decomposition
As in a principal components analysis, if the profiles for a contingency table were plotted in a
multi-dimensional space, they would rarely cover that space in a uniform manner. Rather, the
profiles are often situated geometrically so that most of the variability amongst them is oriented
along a small number of dimensions. A correspondence analysis calculates a set of orthogonal
dimensions such that the first dimension explains as much inertia as possible, the second
explains the next most, etc.
The Inertia and Chi-Square Decomposition displays important information about those
dimensions. The output for the sample data is shown below:
Inertia and Chi-Square Decomposition
Singular Chi- Cumulative
Dimension Value Inertia Square Percentage Percentage Histogram
1 0.1978 0.0391 31.1368 47.1973 47.1973 ***************
2 0.1743 0.0304 24.1831 36.6569 83.8542 ***********
3 0.1043 0.0109 8.6519 13.1146 96.9688 ****
4 0.0501 0.0025 1.9997 3.0312 100.0000 *
TOTAL 0.0829 65.971
This table displays the following information for each dimension:
Singular Value - the square roots of the eigenvalues of a square, symmetric matrix
calculated from the differences between observed and expected profiles.
Inertia - the eigenvalues of that matrix. The largest the inertia for a particular
dimension, the more variability amongst the profiles that dimension represents.
Chi-Square – the contribution of a particular dimension to the chi-squared statistic.
Percentage - the percentage of the total inertia or total chi-square statistic represented
by each dimension.
Cumulative Percentage - the percentage of total inertia represented by a selected
dimension and those extracted earlier. Often, two or three dimensions are sufficient to
represent a large percentage of the total.
Histogram – a graphical representation of each percentage.
In the current example, the first two dimensions account for almost 84% of the total inertia in the
contingency table.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 11
STATGRAPHICS – Rev. 7/6/2009
Scree Plot
A useful plot in helping to determine how many dimensions are necessary to adequately
represent the contingency table is the Scree Plot, which plots the eigenvalues in decreasing
order:
Scree Plot
0.04
0.03
Eigenvalue
0.02
0.01
0
0 1 2 3 4
Dimension
Often, there will be several large values, after which the remainder of the values will be
relatively small.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 12
STATGRAPHICS – Rev. 7/6/2009
Row and Column Contributions
Once the principal dimensions have been calculated, the coordinates of the rows and columns in
those dimensions can be examined. The Row and Column Contributions pane for the sample data
is shown below:
Row and Column Contributions
Row Contributions
Dim #1 Dim #2
Quality Mass Inertia Coord Corr Contr Coord Corr Contr
1 Geology 0.916 0.107 0.137 -0.076 0.055 0.016 -0.303 0.861 0.322
2 Biochemistry 0.881 0.036 0.119 -0.180 0.119 0.030 0.455 0.762 0.248
3 Chemistry 0.644 0.163 0.021 -0.038 0.134 0.006 -0.073 0.510 0.029
4 Zoology 0.929 0.151 0.230 0.327 0.846 0.413 -0.102 0.083 0.052
5 Physics 0.886 0.143 0.196 -0.316 0.880 0.365 -0.027 0.006 0.003
6 Engineering 0.870 0.111 0.152 0.117 0.121 0.039 0.292 0.749 0.310
7 Microbiology 0.680 0.046 0.010 -0.013 0.009 0.000 0.110 0.671 0.018
8 Botany 0.654 0.108 0.067 0.179 0.625 0.088 0.039 0.029 0.005
9 Statistics 0.561 0.036 0.012 -0.125 0.554 0.014 -0.014 0.007 0.000
10 Mathematics 0.319 0.098 0.056 -0.107 0.240 0.029 0.061 0.079 0.012
Supplemental Rows
Dim #1 Dim #2
Quality Mass Inertia Coord Corr Contr Coord Corr Contr
11 Museums 0.556 0.067 0.353 0.314 0.225 0.168 -0.381 0.331 0.318
12 Math Sciences 0.559 0.134 0.041 -0.112 0.493 0.043 0.041 0.066 0.007
Column Contributions
Dim #1 Dim #2
Quality Mass Inertia Coord Corr Contr Coord Corr Contr
1 A 0.587 0.039 0.187 -0.478 0.574 0.228 -0.072 0.013 0.007
2 B 0.816 0.161 0.110 -0.127 0.286 0.067 -0.173 0.531 0.159
3 C 0.465 0.389 0.094 -0.083 0.341 0.068 -0.050 0.124 0.032
4 D 0.968 0.162 0.347 0.390 0.859 0.632 -0.139 0.109 0.103
5 E 0.990 0.249 0.262 0.032 0.012 0.006 0.292 0.978 0.699
Supplemental Columns
Dim #1 Dim #2
Quality Mass Inertia Coord Corr Contr Coord Corr Contr
6 Y 0.807 0.008 0.122 -0.269 0.068 0.014 0.888 0.739 0.196
The output displays:
Quality – a measure of how well a row or column category can be represented by the
number of dimensions that have been extracted relative to its representation using all
dimensions. The closer quality is to 1, the better the representation.
Mass – the proportion of observations in each row and column.
Inertia – the relative inertia of each row and column category. The values sum to 1 for those
categories that were used in the calculations (for rows and columns separately). For
supplemental categories, their size relative to the other categories is of interest.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 13
STATGRAPHICS – Rev. 7/6/2009
Coord. – the principal coordinate or location of the category along the indicated dimension.
These values may be plotted using either the Correspondence Map or the coordinate plots.
Corr. – the correlation between a selected category and the axis defining a given dimension.
Contr. – the relative contribution of an individual category to the inertia of a selected
dimension. The higher the contribution, the more that category contributes to variability
along that dimension.
For example, Zoology and Physics together contribute almost 80% of the column inertia along
dimension #1. On the other hand, the quality of the representation for Mathematics based upon
the two dimensions is relatively low.
2D Correspondence Map
The coordinates of the rows and/or columns may be plotted for any 2 dimensions by selecting the
2D Correspondence Map. By default, the coordinates for the first two dimensions are displayed:
Correspondence Map
rows: principal, columns: principal
1
Y
0.7
Dimension 2
Biochemistry
0.4
E Engineering
0.1 Microbiology
Mathem
Math atics Botany
Sciences
Physics Statistics
A C Chemistry
Zoology
B D
-0.2
Geology
Museums
-0.5
-0.5 -0.2 0.1 0.4 0.7 1
Dimension 1
Different point symbols are used for the rows, columns, and supplementary points.
The distance between any two rows or between any two columns is indicative of their similarity.
For example, note that the points labeled Mathematics, Math Sciences, and Statistics are all very
close together.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 14
STATGRAPHICS – Rev. 7/6/2009
Pane Options
Rows – the type of row coordinates to plot, if any. The weighted average of the squared
principal coordinates equals the eigenvalue of that dimension, while the weighted average of
the squared standard coordinates equals 1.
Columns – the type of column coordinates to plot, if any.
X-Axis Dimension – the dimension to plot along the horizontal axis.
Y-Axis Dimension – the dimension to plot along the vertical axis.
Add reference lines – whether to add vertical and horizontal lines through the origin.
The plot below shows only the columns:
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 15
STATGRAPHICS – Rev. 7/6/2009
Correspondence Map
columns: principal
1
Y
0.7
Dimension 2
0.4
E
0.1
A C
B D
-0.2
-0.5
-0.5 -0.2 0.1 0.4 0.7 1
Dimension 1
Note that the funding levels A, B, C and D line up in sequence along dimension 1. Dimension 2
appears to contrast those 4 levels with E (no funding). The supplemental point Y, for which there
is very little data, is far removed from the other points.
3D Correspondence Map
The coordinates of the rows and/or columns may be plotted for any 3 dimensions by selecting the
3D Correspondence Map. By default, the coordinates for the first three dimensions are
displayed:
Correspondence Map
rows: principal, columns: principal
1 Y
0.7
Dimension 3
Biochemistry
0.4 Mathematics
Math Sciences
E Botany
C Microbiology
0.1 Geology Engineering
Physics Chem istry
B Statistics
Zoology
D
-0.2 A
1
-0.5 Museums 0.7
0.4
-0.5 0.1
-0.2 0.1 0.4 -0.2
0.7 1 -0.5
Dimension 2
Dimension 1
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 16
STATGRAPHICS – Rev. 7/6/2009
Pane Options
Rows – the type of row coordinates to plot, if any.
Columns – the type of column coordinates to plot, if any.
X-Axis Dimension – the dimension to plot along the X axis.
Y-Axis Dimension – the dimension to plot along the Y axis.
Z-Axis Dimension – the dimension to plot along the Z axis.
Add reference lines – whether to add reference lines through the origin.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 17
STATGRAPHICS – Rev. 7/6/2009
Plot of Row Coordinates
This pane displays the row coordinates for each extracted dimension:
Row Coordinate Plot
0.48 Dimension
1
2
Principal coordinate
0.28
0.08
-0.12
-0.32
Geology Chemistry Physics Microbiology Statistics
Biochemistry Zoology Engineering Botany Mathematics
Pane Options
Plot – the type of coordinates to display.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 18
STATGRAPHICS – Rev. 7/6/2009
Plot of Column Coordinates
This pane displays the column coordinates for each extracted dimension:
Column Coordinate Plot
0.52 Dimension
1
0.32 2
Principal coordinate
0.12
-0.08
-0.28
-0.48
A C E
B D
Pane Options
Plot – the type of coordinates to display.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 19
STATGRAPHICS – Rev. 7/6/2009
Save Results
You may save the following results to the datasheet:
1. Point Labels – labels identifying each row and column.
2. Point Quality – the quality of the representation for each row and column.
3. Point Mass – the mass of each row and column.
4. Point Inertia – the relative inertia of each row and column.
5. Principal Coordinates – the principal coordinates for each row and column, for each
extracted dimension.
6. Standard Coordinates – the standard coordinates for each row and column, for each
extracted dimension.
7. Point Correlations – the correlations for each row and column, for each extracted
dimension.
8. Point Contributions - the contributions of each row and column, for each extracted
dimension.
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 20
STATGRAPHICS – Rev. 7/6/2009
Calculations
r = number of rows in contingency table
c = number of columns
v = number of extracted dimensions.
nij = cell count in row i, column j
Totals
c
Row i total: ni. nij (2)
j 1
r
Column j total: n. j nij (3)
i 1
r c
Table total: n.. nij (4)
i 1 j 1
Profiles
nij
Profile for row i: pij (5)
ni .
nij
Profile for column j: qij (6)
n. j
Masses
ni .
Mass for row i: mi . (7)
n..
n. j
Mass for column j: m. j (8)
n..
Expected Frequencies
Expected frequency for cell (i,j): eij n.. mi. m. j (9)
Residuals
Residual for cell (i,j): rij nij eij (10)
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 21
STATGRAPHICS – Rev. 7/6/2009
Chi-Square Distance
rij2
For cell (i,j): d ij (11)
eij
r c
Total: 2 d ij (12)
i 1 j 1
Inertia
d ij
For cell (i,j): inij (13)
n..
2
Total: in.. (14)
n..
Relative Inertia
inij
For cell (i,j): rinij (15)
in..
ini.
For row i: rini. (16)
in..
in. j
For column j: rin. j (17)
in..
Eigenanalysis
Construct r by c matrix T with element:
nij
mi. m. j
n..
t ij (18)
mi. m. j
Calculate eigenvalues eval and eigenvectors evect of matrix
S = T’T (19)
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 22
STATGRAPHICS – Rev. 7/6/2009
Principal Coordinates
c
t
j 1
ij evect jk
For row i, dimension k: rpcik (20)
mi .
evect jk eval k
For column j, dimension k: cpc jk (21)
m. j
Standard Coordinates
rpcik
For row i, dimension k: rscik (22)
eval k
cpc jk
For column j, dimension k: csc jk (23)
eval k
Quality
v
rpc 2
ik
For row i: qi. k 1
(24)
rpc
k
2
ik
cpc 2
jk / rin. j
For column j: q. j k 1
(25)
cpc
k
2
jk / rin. j
Correlation
rpcik2
For row i, dimension k: rcorrik (26)
rpcik2
k
cpc 2jk
For column j, dimension k: ccorr jk (27)
cpc
k
2
jk
Contribution
rpcik2 mi.
For row i, dimension k: rcontri (28)
eval k
cpc 2jk m. j
For column j, dimension k: ccontr j (29)
eval k
2009 by StatPoint Technologies, Inc. Correspondence Analysis - 23
Vous aimerez peut-être aussi
- Provided Data For Exp 6 and Instructions For Data Analysis and Lab ReportDocument2 pagesProvided Data For Exp 6 and Instructions For Data Analysis and Lab ReportFahad Bin HussainPas encore d'évaluation
- Introduction to Statistical Analysis of Laboratory DataD'EverandIntroduction to Statistical Analysis of Laboratory DataPas encore d'évaluation
- Probability & Statistics B: Review of Simple Data SummariesDocument85 pagesProbability & Statistics B: Review of Simple Data SummariesJibreel IsmailPas encore d'évaluation
- 161 - Course Details B.E.computer TechnologyDocument33 pages161 - Course Details B.E.computer Technologyshree_tembePas encore d'évaluation
- Assignment DMBI 2Document2 pagesAssignment DMBI 2IMMORTAL'S PLAYZPas encore d'évaluation
- Epidemiology for Canadian Students: Principles, Methods, and Critical AppraisalD'EverandEpidemiology for Canadian Students: Principles, Methods, and Critical AppraisalPas encore d'évaluation
- Prob-Stat - FinalDocument32 pagesProb-Stat - FinalChú BảyPas encore d'évaluation
- Use of Statistics by ScientistDocument22 pagesUse of Statistics by Scientistvzimak2355Pas encore d'évaluation
- R18B.tech - ECMI IIYearSyllabusDocument52 pagesR18B.tech - ECMI IIYearSyllabusShadow MonarchPas encore d'évaluation
- National Cyber Olympiad: Sample PaperDocument73 pagesNational Cyber Olympiad: Sample PapervsavadiPas encore d'évaluation
- National Cyber Olympiad Sample PaperDocument8 pagesNational Cyber Olympiad Sample PaperpradipkrchakrabartiPas encore d'évaluation
- Exponential Regression Using Microsoft ExcelDocument2 pagesExponential Regression Using Microsoft ExcelCece ChechePas encore d'évaluation
- SyllabusDocument114 pagesSyllabusapi-3748680100% (4)
- Attachment 1 Examination Specification Table (JSU) SPM PhysicsDocument2 pagesAttachment 1 Examination Specification Table (JSU) SPM Physicsthyeoh3383Pas encore d'évaluation
- Computer Science & Engineering-4th Sem 2020-21 B.tech. CSEDocument11 pagesComputer Science & Engineering-4th Sem 2020-21 B.tech. CSEbeastboy00077Pas encore d'évaluation
- Economics Sample Paper Class 11th 23-24 - 240205 - 124435Document5 pagesEconomics Sample Paper Class 11th 23-24 - 240205 - 124435Gauransh DhingPas encore d'évaluation
- Nitc Mechanical Btech Syllabus and CurriculumDocument110 pagesNitc Mechanical Btech Syllabus and Curriculumpikasoc1Pas encore d'évaluation
- Problemas Bono para Tercer Examen de Estadística - Verano 2012Document8 pagesProblemas Bono para Tercer Examen de Estadística - Verano 2012David Meza CarbajalPas encore d'évaluation
- XRDP PATTERN MATCHING PROBABILITY VS IMAGEDocument1 pageXRDP PATTERN MATCHING PROBABILITY VS IMAGEEka Puspa RiniPas encore d'évaluation
- Analysing Geccheriv'cal J LDocument1 pageAnalysing Geccheriv'cal J LChipo ChipPas encore d'évaluation
- Name of Examination-April-May, 2019: Available DetailsDocument6 pagesName of Examination-April-May, 2019: Available DetailsMahesh SinghPas encore d'évaluation
- Final Exam, Data Mining (CEN 871) : Name Surname: Student's IDDocument2 pagesFinal Exam, Data Mining (CEN 871) : Name Surname: Student's IDDimitrios A. KarrasPas encore d'évaluation
- S8EE GradeStat FinalDocument9 pagesS8EE GradeStat FinalVishnu PVPas encore d'évaluation
- STA302 Mid 2010FDocument9 pagesSTA302 Mid 2010FexamkillerPas encore d'évaluation
- DualDegreeCurriculumBTech (CSE) MTech (CSE)Document23 pagesDualDegreeCurriculumBTech (CSE) MTech (CSE)basicelectronic0308Pas encore d'évaluation
- B.Tech Avionics CurriculumDocument97 pagesB.Tech Avionics CurriculumLoves MusicPas encore d'évaluation
- Task 3 memoDocument2 pagesTask 3 memomabotjasefudiPas encore d'évaluation
- Proposed B.Tech ECE Course CurriculumDocument33 pagesProposed B.Tech ECE Course Curriculumrilisab925Pas encore d'évaluation
- 31 BP MQP STATS Question 6Document5 pages31 BP MQP STATS Question 621S1409 -Alden MenezesPas encore d'évaluation
- Edexcel Geography A-Level Fieldwork - Statistical Analysis Techniques NotesDocument11 pagesEdexcel Geography A-Level Fieldwork - Statistical Analysis Techniques NotesSashiPas encore d'évaluation
- Tutorial 3: Statistics With MATLABDocument4 pagesTutorial 3: Statistics With MATLABWahyu Joe PradityoPas encore d'évaluation
- 7 EcDocument38 pages7 EcÅᴅᴀʀsʜ RᴀᴍPas encore d'évaluation
- IAL Biology Statistics GuideDocument22 pagesIAL Biology Statistics GuideAyesha RabbaniPas encore d'évaluation
- Solutions Manual: Ch08-Intro Methods Engrg-S: Review QuestionsDocument5 pagesSolutions Manual: Ch08-Intro Methods Engrg-S: Review QuestionsMahmoud Essam Ahmed100% (1)
- CSBSDocument33 pagesCSBSMadhuPas encore d'évaluation
- Curriculum OF 2017 Undergraduate Admission BatchDocument39 pagesCurriculum OF 2017 Undergraduate Admission BatchDebdutta ChatterjeePas encore d'évaluation
- High-Precision Arithmetic in Mathematical Physics - David H. Bailey - 19 January 2015Document31 pagesHigh-Precision Arithmetic in Mathematical Physics - David H. Bailey - 19 January 2015RMolina65Pas encore d'évaluation
- 161 - Course Details B.E.electrical EngineeringDocument36 pages161 - Course Details B.E.electrical Engineeringkh_chu_1Pas encore d'évaluation
- Logical Analysis of Numerical Data: Mathematical Programming October 2000Document55 pagesLogical Analysis of Numerical Data: Mathematical Programming October 2000KyungMinShimPas encore d'évaluation
- BseeDocument1 pageBseeMigs de LeonPas encore d'évaluation
- B.Tech CSE (DS) - R23Document53 pagesB.Tech CSE (DS) - R23MUKESH KUMARPas encore d'évaluation
- Math 10 Week 8Document6 pagesMath 10 Week 8sushiPas encore d'évaluation
- Mathematics o Level (Form Three) - Statistics - (PDF) EcolebooksDocument26 pagesMathematics o Level (Form Three) - Statistics - (PDF) EcolebooksHANS100% (2)
- Requirements for Minor in Computer Science & EngineeringDocument20 pagesRequirements for Minor in Computer Science & EngineeringAnanyaPas encore d'évaluation
- Organizing and Summarizing Data: StatisticsDocument23 pagesOrganizing and Summarizing Data: Statisticssalah ashrafPas encore d'évaluation
- Parul Practicals 17Document33 pagesParul Practicals 17Himanshi GautamPas encore d'évaluation
- 1 AiDocument58 pages1 AiMajor LoonyPas encore d'évaluation
- B.Tech. Degree Program Course Details and ElectivesDocument47 pagesB.Tech. Degree Program Course Details and ElectivessayakjuPas encore d'évaluation
- Unit: 5-Curve Fitting by Numerical MethodsDocument34 pagesUnit: 5-Curve Fitting by Numerical MethodsBHAVESHPas encore d'évaluation
- Gage Linearity and Accuracy AnalysisDocument10 pagesGage Linearity and Accuracy AnalysisRamdas PaithankarPas encore d'évaluation
- Simple Pendulum: Experiment ADocument6 pagesSimple Pendulum: Experiment AEmmanuelPas encore d'évaluation
- Tutorial Sheet 1Document4 pagesTutorial Sheet 1Devang SinghPas encore d'évaluation
- MIT Feedback Systems Final Exam SolutionsDocument11 pagesMIT Feedback Systems Final Exam SolutionsLuis AntonioPas encore d'évaluation
- Pre Board XII CS 21 22 1Document6 pagesPre Board XII CS 21 22 1aleksePas encore d'évaluation
- BTech CSEDocument28 pagesBTech CSEKaushalPranavPas encore d'évaluation
- STAT 342 Statistical Methods For Engineers: Homework#01Document2 pagesSTAT 342 Statistical Methods For Engineers: Homework#01HamzaBaigPas encore d'évaluation
- Av2012 FinalDocument52 pagesAv2012 FinalabhishekagarwlPas encore d'évaluation
- Sampling DistributionsDocument3 pagesSampling DistributionsAnonymous FZNn6rBPas encore d'évaluation
- Dashboard GageDocument4 pagesDashboard GageAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Screening DesignsDocument50 pagesDOE Wizard - Screening DesignsAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Process and Mixture FactorsDocument46 pagesDOE Wizard - Process and Mixture FactorsAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Quantitative and Categorical FactorsDocument44 pagesDOE Wizard - Quantitative and Categorical FactorsAnonymous FZNn6rB100% (1)
- DOE Wizard - Response Surface DesignsDocument24 pagesDOE Wizard - Response Surface DesignsAnonymous FZNn6rBPas encore d'évaluation
- Quantile PlotDocument4 pagesQuantile PlotAnonymous FZNn6rBPas encore d'évaluation
- Data ViewerDocument3 pagesData ViewerAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Robust Parameter DesignsDocument20 pagesDOE Wizard - Robust Parameter DesignsAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Single Factor Categorical DesignsDocument19 pagesDOE Wizard - Single Factor Categorical DesignsAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Variance Component DesignsDocument14 pagesDOE Wizard - Variance Component DesignsAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Mixture DesignsDocument34 pagesDOE Wizard - Mixture DesignsAnonymous FZNn6rBPas encore d'évaluation
- Multiple Correspondence AnalysisDocument16 pagesMultiple Correspondence AnalysisAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Multi-Factor Categorical DesignsDocument17 pagesDOE Wizard - Multi-Factor Categorical DesignsAnonymous FZNn6rBPas encore d'évaluation
- Design of Experiments WizardDocument70 pagesDesign of Experiments WizardAnonymous FZNn6rBPas encore d'évaluation
- DOE Wizard - Multilevel Factorial DesignsDocument11 pagesDOE Wizard - Multilevel Factorial DesignsAnonymous FZNn6rBPas encore d'évaluation
- Monte Carlo Simulation (ARIMA Time Series Models)Document15 pagesMonte Carlo Simulation (ARIMA Time Series Models)Anonymous FZNn6rBPas encore d'évaluation
- Sequential Sampling: Sample MeasurementDocument14 pagesSequential Sampling: Sample MeasurementAnonymous FZNn6rB100% (1)
- Sample Size Determination (Capability Indices)Document4 pagesSample Size Determination (Capability Indices)Anonymous FZNn6rBPas encore d'évaluation
- Andrews PlotDocument5 pagesAndrews PlotAnonymous FZNn6rBPas encore d'évaluation
- Monte Carlo Simulation (General Simulation Models)Document11 pagesMonte Carlo Simulation (General Simulation Models)Anonymous FZNn6rBPas encore d'évaluation
- Statistical Tolerance Limits (Observations)Document11 pagesStatistical Tolerance Limits (Observations)Anonymous FZNn6rBPas encore d'évaluation
- Parallel Coordinates PlotDocument5 pagesParallel Coordinates PlotAnonymous FZNn6rBPas encore d'évaluation
- Chernoff FacesDocument8 pagesChernoff FacesAnonymous FZNn6rBPas encore d'évaluation
- Star Glyphs and Sunray PlotsDocument6 pagesStar Glyphs and Sunray PlotsAnonymous FZNn6rBPas encore d'évaluation
- Monte Carlo Simulation (Random Number Generation)Document9 pagesMonte Carlo Simulation (Random Number Generation)Anonymous FZNn6rBPas encore d'évaluation
- Sample Size Determination (Tolerance Limits)Document6 pagesSample Size Determination (Tolerance Limits)Anonymous FZNn6rBPas encore d'évaluation
- Joe Hisaishi Castle in The Sky (Innocent)Document2 pagesJoe Hisaishi Castle in The Sky (Innocent)LenWez100% (4)
- Adams 2 Nast Ran TestsDocument26 pagesAdams 2 Nast Ran TestsAnonymous FZNn6rBPas encore d'évaluation
- SONY IPELA g50 Operation Manual v2-1 enDocument305 pagesSONY IPELA g50 Operation Manual v2-1 enDoruSPas encore d'évaluation
- Event Planning Rubric - Final Project Luc PatbergDocument3 pagesEvent Planning Rubric - Final Project Luc Patbergapi-473891068Pas encore d'évaluation
- Andrew Nathan Lee MR Tp059923 Fyp Apu3f2209 CsdaDocument297 pagesAndrew Nathan Lee MR Tp059923 Fyp Apu3f2209 CsdaHema Latha Krishna NairPas encore d'évaluation
- Make Way For The Algorithms Symbolic Actions and Change in A Regime of KnowingDocument26 pagesMake Way For The Algorithms Symbolic Actions and Change in A Regime of KnowingAbimbola PotterPas encore d'évaluation
- Irobot AccessoriesDocument24 pagesIrobot AccessoriesPredatorBDU.com100% (1)
- Mercantile - 13 05 2021Document1 pageMercantile - 13 05 2021AlexMason100% (1)
- Bulk Carrier Safety ABS RulesDocument3 pagesBulk Carrier Safety ABS RulesHUNG LE THANHPas encore d'évaluation
- Qlogic SAN CLI GuideDocument408 pagesQlogic SAN CLI Guide김대성Pas encore d'évaluation
- Tripp Lite Owners Manual 798383 PDFDocument32 pagesTripp Lite Owners Manual 798383 PDFMathias VillasecaPas encore d'évaluation
- ServiceNow Certified System Administrator CSA Practice Test Set 18Document151 pagesServiceNow Certified System Administrator CSA Practice Test Set 18Apoorv DiwanPas encore d'évaluation
- Ricoh Printer 3245C-Troubleshooting ManualDocument82 pagesRicoh Printer 3245C-Troubleshooting ManualPeter xuPas encore d'évaluation
- Chapter 6 - MS PowerPoint Advance FeatureDocument41 pagesChapter 6 - MS PowerPoint Advance FeaturePhrexilyn PajarilloPas encore d'évaluation
- Effective Use of Instagram For BusinessDocument10 pagesEffective Use of Instagram For BusinessDemand MetricPas encore d'évaluation
- Chargeback of Refund in My CreditcardDocument2 pagesChargeback of Refund in My CreditcardtdxdvnddmfPas encore d'évaluation
- Fungsi Komposisi English YOLANDADocument11 pagesFungsi Komposisi English YOLANDAyolanda anastasyaPas encore d'évaluation
- E7756v1 3 PDFDocument82 pagesE7756v1 3 PDFxmiePas encore d'évaluation
- Midiengine 8Port/Se™ User'S Manual: UsicquestDocument66 pagesMidiengine 8Port/Se™ User'S Manual: UsicquestNuno FilipePas encore d'évaluation
- Struggles of ICT Students Who LackDocument13 pagesStruggles of ICT Students Who LackAjel Rafael Buenviaje100% (1)
- Employee Roster with Company, Name, TitleDocument12 pagesEmployee Roster with Company, Name, TitleMahesh MalvePas encore d'évaluation
- Computer 4 - 1st Monthly TestDocument3 pagesComputer 4 - 1st Monthly TestMaria JocosaPas encore d'évaluation
- PHD Thesis Commerce PDFDocument7 pagesPHD Thesis Commerce PDFufagmcgld100% (1)
- GSM Training Customer Care InductionDocument63 pagesGSM Training Customer Care InductionfirasibraheemPas encore d'évaluation
- COMMANDER 1900 Series Circular Chart Recorders: Programming GuideDocument60 pagesCOMMANDER 1900 Series Circular Chart Recorders: Programming Guideefl321Pas encore d'évaluation
- Communicate with embedded devices via Bluetooth with the ZX-BLUETOOTH boardDocument10 pagesCommunicate with embedded devices via Bluetooth with the ZX-BLUETOOTH boardashwanisingla013Pas encore d'évaluation
- Singular Value Array AND: Reducing The Computations of The Decomposition Given by Brent Luk YangDocument13 pagesSingular Value Array AND: Reducing The Computations of The Decomposition Given by Brent Luk YangDeepthi Priya BejjamPas encore d'évaluation
- CMP Catalogue 2014Document44 pagesCMP Catalogue 2014Joseph TingPas encore d'évaluation
- Mahesh Kumar GS - Technical Project Manager - Technical Lead - Senior Developer - DeveloperDocument9 pagesMahesh Kumar GS - Technical Project Manager - Technical Lead - Senior Developer - Developernagaraj.hPas encore d'évaluation
- Optimal Drug Dosage Control Strategy of Immune Systems Using Reinforcement LearningDocument10 pagesOptimal Drug Dosage Control Strategy of Immune Systems Using Reinforcement LearninggeethakaniPas encore d'évaluation
- 19XR, XRV Product Data PDFDocument56 pages19XR, XRV Product Data PDFCristian Ramos PPas encore d'évaluation
- Contact Session 2Document35 pagesContact Session 2api-3724082Pas encore d'évaluation