Académique Documents
Professionnel Documents
Culture Documents
Dinosaurs Before Dark
Transféré par
Ina Simache0 évaluation0% ont trouvé ce document utile (0 vote)
30 vues9 pagesMolecular diagnostics is increasingly important in cancer care for diagnosis, prognosis, predicting therapy response, and monitoring minimal residual disease. Molecular biomarkers include genetic and epigenetic abnormalities that can be detected through various methods like chromosomal translocations, gene expression changes, and mutations. Key applications include detecting translocations for diagnosing cancers like CML and Burkitt lymphoma, assessing clonality in uncertain lymphoid proliferations, and identifying amplifications like MYCN that predict poor outcomes in neuroblastoma. Molecular diagnostics provides critical information to guide patient care and treatment decisions.
Description originale:
bsgdhbsnb
Copyright
© © All Rights Reserved
Formats disponibles
DOCX, PDF, TXT ou lisez en ligne sur Scribd
Partager ce document
Partager ou intégrer le document
Avez-vous trouvé ce document utile ?
Ce contenu est-il inapproprié ?
Signaler ce documentMolecular diagnostics is increasingly important in cancer care for diagnosis, prognosis, predicting therapy response, and monitoring minimal residual disease. Molecular biomarkers include genetic and epigenetic abnormalities that can be detected through various methods like chromosomal translocations, gene expression changes, and mutations. Key applications include detecting translocations for diagnosing cancers like CML and Burkitt lymphoma, assessing clonality in uncertain lymphoid proliferations, and identifying amplifications like MYCN that predict poor outcomes in neuroblastoma. Molecular diagnostics provides critical information to guide patient care and treatment decisions.
Droits d'auteur :
© All Rights Reserved
Formats disponibles
Téléchargez comme DOCX, PDF, TXT ou lisez en ligne sur Scribd
0 évaluation0% ont trouvé ce document utile (0 vote)
30 vues9 pagesDinosaurs Before Dark
Transféré par
Ina SimacheMolecular diagnostics is increasingly important in cancer care for diagnosis, prognosis, predicting therapy response, and monitoring minimal residual disease. Molecular biomarkers include genetic and epigenetic abnormalities that can be detected through various methods like chromosomal translocations, gene expression changes, and mutations. Key applications include detecting translocations for diagnosing cancers like CML and Burkitt lymphoma, assessing clonality in uncertain lymphoid proliferations, and identifying amplifications like MYCN that predict poor outcomes in neuroblastoma. Molecular diagnostics provides critical information to guide patient care and treatment decisions.
Droits d'auteur :
© All Rights Reserved
Formats disponibles
Téléchargez comme DOCX, PDF, TXT ou lisez en ligne sur Scribd
Vous êtes sur la page 1sur 9
3.
Molecular Methods in Cancer
Applications of Molecular Diagnostics in Oncology
Molecular diagnostics is increasingly impacting a number of areas of cancer care delivery including
diagnosis, prognosis, in predicting response to particular therapies, and in minimal residual disease
monitoring.
Each of these depends on detection or measurement of one or more disease-specific molecular
biomarkers representing abnormalities in genetic or epigenetic pathways controlling cellular proliferation,
differentiation, or cell death (Table 3.1). In addition, molecular diagnostics is beginning to play a role in
predicting host metabolism of drugs—for example, in predicting fast versus slow thiopurine metabolizers
using polymorphisms in the thiopurine methyltransferase (TPMT) allele and in use in dosing patients with
thiopurine drugs.
Molecular diagnostics has also had a major impact on assessing an engraftment after bone marrow
transplantation and in tissue typing for bone marrow and solid organ transplantation.
The ideal cancer biomarker is only associated with the disease and not the normal state. The utility of
the biomarker largely depends on what the clinical effect the biomarker predicts for, how large the effect is,
and how strong the evidence is for the effect.
For clinical application, biomarkers need a high level of analytic validity, clinical validity, and clinical
utility. Analytic validity refers to the ability of the overall testing process to accurately detect and, in many
cases, measure the biomarker. Clinical validity is the ability of a biomarker to predict a particular disease
behavior or response to therapy. Clinical utility, arguably the most difficult to assess, addresses whether the
information available from the biomarker is actually beneficial for patient care.
Biomarkers can take many forms including chromosomal translocations and other chromosomal
rearrangements, gene amplification, copy number variation, point mutations, single nucleotide
polymorphisms, changes in gene expression (including micro RNAs), and epigenetic alterations. Most
biomarkers in widespread use represent either gain of function or loss of function alterations in key signaling
pathways. Those that occur early and at a high frequency in tumors tend to be driver mutations, whose
function is important for the cancer cell’s proliferation and/or survival. These are particularly useful as
biomarkers because they often representimportant therapeutic targets. However, cancer cells accumulate
many genetic alterations, called passenger mutations, which tend to occur at a lower frequency overall and in
a subset of a heterogeneous population of tumor cells that may contribute to the cancer phenotype but are
not absolutely essential. Distinguishing passenger from driver mutations using various functional assays has
become a major focus of translational research in cancer. The same biomarker may have utility in a variety of
settings. For example, the detection of the BCR-ABL1 translocation, pathognomonic for chronic myelogenous
leukemia (CML), is used for establishing the diagnosis, for the selection of therapy, and for monitoring for
minimal residual disease during and after therapy.
Some of the most heavily used genetic biomarkers in cancer, particularly in hematologic malignancies,
are chromosomal translocations.
For certain diseases such as CML, detection of the BCR-ABL1 translocation or in Burkitt lymphoma the
immunoglobulin gene-MYC translocation is required, according to current World Health Organization (WHO)
guidelines, to make the diagnosis.
Identification of translocations is important in the diagnosis and subtyping of acute leukemias (e.g.,
detection of PML-RARA and variant translocations in acute promyelocytic leukemia) and is also extremely
important for the diagnosis of sarcomas such as Ewing sarcoma. The discovery of chromosomal translocations,
such as the TMPRSS-ETS in prostate cancer and ALK translocations in non– small-cell lung cancer, portends an
importance of detecting translocations in solid tumors. Chromosomal translocations, especially for
hematologic malignancies, have been traditionally detected by classical karyotyping. This approach has
limitations; in particular, it requires viable, dividing cells, which are often not readily available from solid
tumor biopsies. In addition, a significant proportion of chromosomal translocations are not detectable by
conventional karyotyping. For example, 5% to 10% of CML cases lack detectable t(9;22) by G banding. Such
“cryptic” translocations require other approaches for detection, which are to be discussed, including
fluorescent in situ hybridization (FISH), polymerase chain reaction (PCR), as well as nucleic acid sequencing-
based methods.
In certain settings, it can be helpful to detect if a population of cells is clonal. For example, in some
lymphoid infiltrates, the cells are well differentiated and it can be difficult to determine whether these
represent a reactive or neoplastic infiltrate. If dispersed, cells are available and these could be analyzed by
flow cytometer to detect whether a monotypic population expressing either immunoglobulin kappa or lambda
light chains is present. In theory, immunohistochemical staining (IHC) for immunoglobulin light chains could be
used to assess clonality; however, in practice this is done with more sensitivity using RNA in situ hybridization
for immunoglobulin kappa and lambda light chain transcripts. The most sensitive way to detect clonality in a
B-cell population is to analyze the size of the break point cluster region that arises as a result of VDJ
recombination by PCR. Reactive B cells will show a distribution in the size of the VDJ recombination for the IGH
or IGK or IGL, whereas clonal cells will show a predominant band that represents the size of the VDJ region of
the dominant clone.
Similarly, sometimes it can be difficult to distinguish neoplastic from reactive T-cell infiltrates. Given
the large number of T-cell antigen receptors, it is not as simple to detect clonality by IHC or flow cytometry in
T-cell proliferations. One approach is to use aberrant loss of T-cell antigen expression to aid in the diagnosis of
T-cell neoplasms. Another is to detect clonal rearrangement of the VDJ region of the T-cell receptor gamma
(TCR ) gene, which can be done by PCR on both fresh and formalin-fixed paraffinembedded (FFPE) tissue.
Gene amplification is another important mechanism in cancer that has been found to have high utility
in a subset of cancers.
MYCN amplification occurs in approximately 40% of undifferentiated or poorly differentiated
neuroblastoma subtypes, either appearing as double minute chromosomes or homogeneously of poor
outcomes, particularly in patients with localized (stage 1 or stage 2) disease or in infants with stage 4S
metastatic disease, where fewer than half of patients survive beyond 5 years.
Use of other chromosome abnormalities has been largely limited to the diagnosis and prognostication
of hematologic disorders.
Roughly half of all myelodysplastic disorders show cytogenetically detectable chromosomal
abnormalities, such as monosomy 5 or 7, partial chromosomal loss (5q-, 7q-), or complex chromosomal
abnormalities. Certain abnormalities in isolation (e.g., 5q-) have a favorable prognosis, whereas many others
(e.g., “complex” karyotypes with three or more abnormalities) carry a worse prognosis.
Differences in ploidy have proven to be useful predictors in pediatric acute lymphocytic leukemia (ALL),
with hyperdiploid cases (>50 chromosomes) showing a distinctly more favorable course compared with
hypodiploid or near diploid cases. Overall, DNA ploidy can be assessed by flow cytometry. Specific
chromosomal copy number alterations can be detected by conventional karyotyping, array hybridization
methods, or FISH.
Copy number variation (CNV) represents the most common type of structural chromosomal alteration.
Regions affected by CNVs range from approximately 1 kilobase to several megabases that are either amplified
or deleted. It is estimated that about 0.4% of the genomes of healthy individuals differ in copy number. CNVs
resulting in deletion of genes such as BRCA1, BRCA2, APC, mismatch repair genes, and TP53 have been
implicated in a wide range of highly penetrant cancers. CNVs can be detected by a variety of means including
FISH, comparative or array genomic hybridization, or virtual karyotyping using single nucleotide polymorphism
(SNP) arrays.
Increasingly, CNV is detected using next-generation sequencing. Large-scale sequencing of tumors has
identified many mutations that are of potential prognostic and therapeutic significance.
As will be discussed further, a wide range of strategies is available for the detection of point mutations (Fig.
3.1). It is important to recognize that many nucleotide variations occur at any given allele in populations.
Formally, the term polymorphism is used to describe genetic differences present in ≥1% of the human
population, whereas mutation describes less frequent differences. However, in practice, polymorphism is
often used to describe a nonpathogenic genetic change, and mutation a deleterious change, regardless of
their frequencies.
Mutations can be classified according to their effect in the structure of a gene. The most common of
these disease-associated alterations are single nucleotide substitutions (point mutations); however, many
deletions, insertions, gene rearrangements, gene amplification, and copy number variations have been
identified that have clinical significance. Point mutations may affect promoters, splicing sites, or coding
regions.
Coding region mutations can be classified into three kinds, depending on the impact on the codon:
missense mutation, a nucleotide change leads to the substitution of an amino acid to another; nonsense
mutation, a nucleotide substitution causes premature termination of codons with protein truncation; and
silent mutation, a nucleotide change does not change the coded amino acid.
Loss of function mutations, either through point mutations or deletions in tumor suppression genes
such as APC and TP53, are the most common mutations in cancers. Tumor suppression genes require two-hit
(biallelic) mutations that inactivate both copies of the gene in order to allow tumorigenesis to occur. The first
hit is usually an inherited or somatic point mutation, and the second hit is assumed to be an acquired deletion
mutation that deletes the second copy of the tumor suppression gene. Promoter methylation of tumor
suppressor genes is an alternative route to tumorigenesis that, to date, has not been commonly employed for
molecular diagnostics.
Oncogenes originate from the deregulation of genes that normally encode for proteins associated with
cell growth, differentiation, apoptosis, and signal transduction (proto-oncogenes, [e.g., BRAF and KRAS]).
Proto-oncogenes generally require only one gain of function or activating mutation to become oncogenic.
Common mutation types that result in proto-oncogene activation include point mutations, gene
amplifications, and chromosomal translocations. One example is mutations in the epidermal growth factor
receptor (EGFR) that occur in lung cancer, which are almost exclusively seen in nonmucinous bronchoalveolar
carcinomas. Somatic mutations of EGFR constitutively activate the receptor tyrosine kinase (TK). Importantly,
responsiveness of tumors harboring these mutations to the inhibitor gefitinib is highly coordinated with a
mutation of the EGFR TK domain.
One of the challenges with using mutations as biomarkers is that there can be many nucleotide
alterations that affect a given gene. For example, there are over 100 known different point mutations in EGFR
reported in non–small-cell lung cancer. Many of these mutations occur at low frequency and have an
unknown clinical significance. Another important concept is that the same driver oncogene may be mutated in
a variety of different tumors. For example, lung cancers harbor a number of other different alterations that
are common in other solid tumors, which generally occur at lower frequencies than EGFR mutations such as
KRAS, BRAF, and HER2. Some lung cancers have translocations involving the ALK kinase gene. ALK,
interestingly, is also activated by point mutations in a neuroblastoma as by translocation in anaplastic large
cell lymphoma (Fig. 3.2). Hence, a therapy targeted to a genetic alteration in one cancer may demonstrate
efficacy in other cancers.
The detection of mutations is also important in the evaluation of chemotherapy resistance. Roughly a
third of CML patients are resistant to the frontline ABL1 kinase inhibitor imatinib, either at the time of initial
treatment or, more commonly, secondarily.
In cases of primary failure or secondary failure, over 100 different ABL1 mutations have been
identified, including particularly common ones such as T315I and P loop mutations. While some mutations,
such as Y253H, respond to second generation TK inhibitors (TKI), others, such as the T315I mutation, are
noteworthy because they confer resistance not only to imatinib, but also to nilotinib and dasatinib.
Mutations are also used as important predictive biomarkers (Table 3.1). Two of the most notable
examples are the use of the BRCA1 and BRCA2 mutation analysis for women with a strong family history of
breast cancer. Over 200 mutations (loss of function point mutations, small deletions, or insertions) occur in
BRCA genes, which are distributed across the genes necessitating full sequencing for their detection. The
overall prevalence of these occur in about 0.1% of the general population.
The lifetime risk of breast cancer for women carrying BRCA1 mutations is in the range of 47% to 66%,
whereas for BRCA2 mutations, it is in the range of 40% to 57%. In addition, the risk of other tumors including
ovarian, fallopian, and pancreatic cancer is also increased. Detection of BRCA1 and BRCA2 mutations is,
therefore, important for cancer prevention and risk reduction.
The Clinical Molecular Diagnostics. Laboratory: Rules and Regulations
Laboratories in the United States that perform molecular diagnostic testing are categorized as high-
complexity laboratories under the Clinical Laboratory Improvement Amendments of 1988 (CLIA). The CLIA
program sets the minimum administrative and technical standards that must be met in order to ensure quality
laboratory testing. Most laboratories in the United States that perform clinical testing in humans are regulated
under CLIA. CLIA-certified laboratories must be accredited by professional organizations such as the Joint
Commission, the College of American Pathologists, or another agency officially approved by the Centers for
Medicare & Medicaid Services (CMS), and must comply with CLIA standards and guidelines for quality
assurance. Although the regulation of laboratory services is in the U.S. Food and Drug Administration’s (FDA)
jurisdiction, the FDA has historically exercised enforcement discretion. Therefore, FDA approval is not
currently required for clinical implementation of molecular tests as long as other regulations are met.
Specimen Requirements for Molecular Diagnostics
Samples typically received for molecular oncology testing include blood, bone marrow aspirates and
biopsies, fluids, organ-specific fresh tissues in saline or tissue culture media such as Roswell Park Memorial
Institute (RPMI), FFPE tissues, and cytology cell blocks. Molecular tests can be ordered electronically or
through written requisition forms, but never through verbal requests only.
All samples submitted for molecular testing need to be appropriately identified. Sample type, quantity,
and specimen handling and transport requirements should conform to the laboratory’s stated requirements in
order to ensure valid test results.
Blood and bone marrow samples should be drawn into anticoagulated tubes. The preferred
anticoagulant for most molecular assays is ethylenediaminetetraacetic acid (EDTA; lavender). Other
acceptable collection tubes include ACD (yellow) solutions A and B. Heparinized tubes are not preferred for
most molecular tests because heparin inhibits the polymerase enzyme utilized in PCR, which may lead to assay
failure. Blood and bone marrow samples can be transported at ambient temperature. Blood samples should
never be frozen prior to separation of cellular elements because this causes hemolysis, which interferes with
DNA amplification. Fluids should be transported on ice. Tissues should be frozen (preferred method) as soon
as possible and sent on dry ice to minimize degradation. Fresh tissues in RPMI should be sent on ice or cold
packs. Cells should be kept frozen and sent on dry ice; DNA samples can be sent at ambient temperature or on
ice.
For FFPE tissue blocks, typical collection and handling procedures include cutting 4 to 6 microtome
sections of 10-micron thickness each on uncoated slides, air-drying unstained sections at room temperature,
and staining one of the slides with hematoxylin and eosin (H&E). A board-certified pathologist reviews the
H&E slides to ensure the tissue block contains a sufficient quantity of neoplastic tumor cells, and circles an
area on the H&E slide that will be used as a template to guide macrodissection or microdissection of the
adjacent, unstained slides. The pathologist also provides an estimate of the percentage of neoplastic cells in
the area that will be tested, which should exceed the established limit of detection (LOD) of the assay.
Molecular Diagnostics Testing Process
The workflow of a molecular test begins with receipt and accessioning of the specimen in the clinical
molecular diagnostics laboratory followed by extraction of the nucleic acid (DNA or RNA), test setup, detection
of analyte (e.g., PCR products), data analysis, and result reporting to the patient medical record (Fig. 3.3).
An extraction of intact, moderately high-quality DNA is essential for molecular assays. For DNA
extraction, the preferred age for blood, bone marrow, and fluid samples is less than 5 days; for frozen or fixed
tissue, it is indefinite; and for fresh tissue, it is overnight.
Although there is no age limit for the use of a fixed and embedded tissue specimen for analysis, older
specimens may yield a lower quantity and quality of DNA. Because RNA is significantly more labile than DNA,
the preferred age for blood and bone marrow is less than 48 hours (from time of collection). Tissue samples
intended for an RNA analysis should be promptly processed in fresh state, snap frozen, or preserved with RNA
stabilizing agents for transport.
Dedicated areas, equipment, and materials are designated for various stages of DNA and RNA
extraction procedures. DNA and RNA isolation can be done by manual or automated methods.
Currently, most clinical laboratories employ commercial protocols based on liquid- or solid-phase
extractions. Nucleated cells are isolated from biological samples prior to nucleic acid extraction.
White blood cells (WBC) can be isolated from blood and bone marrow samples by different methods.
One method involves lysing the red blood cells with an ammonium chloride solution, which yields the total
WBC population and other nucleated cells present.
Another method involves a gradient preparation with a Ficoll solution, which yields the mononuclear
cell population only. Sections of FFPE tissue blocks are prepared for DNA extraction by first removing the
paraffin and disrupting the cell membranes with proteinase K digestion. Fresh and frozen tissues also undergo
proteinase K digestion prior to nucleic acid extraction. DNA isolation protocols consist of several steps,
including cell lysis, DNA purification by salting out the proteins and other debris (nonorganic method), or by
solvent extractions of the proteins with phenol and chloroform solutions (organic method). The DNA is then
precipitated out of the solution with isopropanol or ethanol. The pellet is washed with 70% to 80% ethanol
and then solubilized in buffer, such as Tris-EDTA solution. Proteinase K can be added to assist in the disruption
and to prevent nonspecific degradation of the DNA. RNase is sometimes added to eliminate contaminating
RNA. The DNA yield is quantitated spectrophotometrically, and the DNA sample integrity is visually checked, if
necessary, on an agarose gel followed by ethidium bromide staining. Intact DNA appears as a high–molecular-
weight single band, whereas degraded DNA is identified as a smear of variably sized fragments. After
extraction, the DNA is stored at 4°C prior to use in a PCR assay, and is then stored at –70°C after completion of
the assay. Because the DNA extracted from formalin-fixed tissue is degraded to a variable extent, an analysis
of the extraction product by gel electrophoresis is not informative. Yield and integrity of the extracted DNA is
best assessed by an amplification control to ensure that the quality and quantity of input DNA is adequate to
yield a valid result.
RNA isolation steps are similar to the ones described previously for DNA extraction. However, RNA is
inherently less stable than DNA due to its single-strand conformation and susceptibility to degradation by
RNase, which is ubiquitous in the environment.
To ensure preservation of target RNA, special precautions are required, including the use of
diethylpyrocarbonate (DEPC) water in all reagents used in RNA procedures, and special decontamination of
work area and pipettes to prevent RNase contamination. The extracted RNA is usually degraded to a variable
extent so that the analysis of the extraction product by gel electrophoresis is not informative.
The quality of the RNA and its suitability for use in a reverse transcriptase polymerase chain reaction
(RT-PCR)–based assay is assessed most appropriately by the demonstration of a positive result in an assay
designed to detect the RNA transcripts for a “housekeeping gene,” such as ABL1 or GAPDH. Any RNA sample in
which the 260/280-nm absorption ratio is below 1.9 or greater than 2.0 may contain contaminants and must
be cleaned prior to analysis.
Following nucleic acid extraction, the assay is set up according to written procedures established
during validation/verification of the assay by qualified laboratory staff. Dedicated areas, equipment, and
materials are designated for various stages of the test (e.g., extraction, pre-PCR and post-PCR for
amplification-based assays).
For each molecular oncology test, appropriated positive and negative control specimens are included
to each run as a matter of routine quality assessment. A no template (blank) control, containing the complete
reaction mixture except for nucleic acids, is also included in amplification-based assays to evaluate for
amplicon contamination in the assay reagents that may lead to inaccurate results. The controls are processed
in the same manner as patient samples to ensure that established performance characteristics are being met
for each step of the assay (extraction, amplification, and detection). All assay controls and overall performance
of the run must be examined prior to interpretation of sample results. Following acceptance of the controls,
results are electronically entered into reports. The final report is reviewed and signed by the laboratory
director or a qualified designee who meets the same qualifications as the director, as defined by CLIA (see
previous).
Technologies
Several traditional and emerging techniques are currently available for mutation detection in cancer
(Table 3.2). In the era of personalized medicine, molecular oncology assays are rapidly moving from a
mutational analysis of single genes toward a multigene panel analysis. As the number of “actionable”
mutations such as ALK, EGFR, BRAF, and others increase, the use of next-generation sequencing platforms is
expected to become much more widespread.
Both traditional and emerging testing approaches have advantages and disadvantages that need to be
balanced before a test platform is implemented into practice.
An important consideration when adding a new oncology test in the clinical laboratory menu is to
define the intended use of the assay (e.g., diagnosis, prognosis, prediction of therapy response).
The clinical utility of the assay, appropriate types of specimens, the spectrum of possible mutations
that can be found in the genomic region of interest, and available methods for testing should also be
determined. The laboratory director and ordering physicians should also discuss the estimated test volume,
optimal reporting format, and required turnaround time for the proposed new test.
Polymerase Chain Reaction
Polymerase chain reaction (PCR) is widely used in all molecular diagnostics laboratories for the rapid
amplification of targeted DNA sequences. The reaction includes the specimen template DNA, forward and
reverse primers (18 to 24 oligonucleotides long), Taq DNA polymerase, and each of the four nucleotides bases
(dATP, dTTP, dCTP, dGTP). During PCR, selected genomic sequences undergo repetitive temperature cycling
(sequential heat and cooling) that allows for denaturation of double-stranded DNA template, annealing of the
primers to the targeted complementary sequences on the template, and extension of new strands of DNA by
Taq polymerase from nucleotides, using the primers as the starting point. Each cycle doubles the copy number
of PCR templates for the next round of polymerase activity, resulting in an exponential amplification of the
selected target sequence. The PCR products (amplicons) are detected by electrophoresis or in real-time
systems simultaneously to the amplification reaction (see real-time PCR, which follows).
PCR is specifically designed to work on DNA templates because the Taq polymerase does not recognize
RNA as a starting material. Nonetheless, PCR can be adapted to RNA testing by including a reverse
transcription step to convert a RNA sequence into its cognate cDNA sequence before the PCR reaction is
performed (see reverse-transcription PCR, which follows). Multiplex PCR reactions can also be designed with
multiple primers for simultaneous amplification of multiple genomic targets. PCR is a highly sensitive and
specific technique that can be employed in different capacities for the detection of point mutations, small
deletions, insertions and duplications, as well as gene rearrangements and clonality assessment. Limits of
detection can reach 0.1% mutant allele or lower, which is important for the detection of somatic mutations in
oncology because tumor specimens are usually composed of a mixture of tumor and normal cells. Reverse
transcription PCR can also be used for the relative quantification of target RNA in minimal residual disease
testing, such as BCRABL1 transcripts in CML. Another advantage of PCR is its ability to amplify small amounts
of low quality FFPE-derived DNA. However, applications of PCR can be limited because it cannot amplify across
large or highly repetitive genomic regions. Also, the PCR reaction can be inhibited by heparin or melanin if
present in the extracted DNA, which may lead to assay failure. Finally, the risk of false positives due to
specimen or amplicon contamination is an important issue when using PCR-based techniques; therefore,
stringent laboratory procedures, as described previously, are used to minimize contamination. With the
exception of hybridization assays, such as fluorescence in situ hybridization and genomic microarrays,
PCR is the necessary initial step in all current molecular oncology assays.
Targeted Mutation Analysis Methods
Real-Time PCR (q-PCR)
In real-time PCR (q-PCR), the polymerase chain reaction is performed with a PCR reporter that is
usually a fluorescent doublestranded DNA binding dye or a fluorescent reporter probe. The intensity of the
fluorescence produced at each amplification cycle is monitored in real time, and both quantification and
detection of targeted sequences is accomplished in the reaction tube as the PCR amplification proceeds.
The intensity of the fluorescent signal for a given DNA fragment (wild type or mutant) is correlated
with its quantity, based on the PCR cycle in which the fluorescence rises above the background (crossing
threshold [Ct] or crossing point [Cp]). The Ct value can be used for qualitative or quantitative analysis.
Qualitative assays use the Ct as a cutoff for determining “presence” or “absence” of a given target in the
reaction. A qualitative analysis by q-PCR is particularly useful for a targeted detection of point mutations
that are located in mutational hotspots. Examples include the JAK2 V617F mutation, which is located within
exon 14, and is found in several myeloproliferative neoplasms (polycythemia vera, essential
thrombocythemia, and primary myelofibrosis), and the BRAF V600E, which is located within exon 15, and is
found in various cancer types including melanomas and thyroid and lung cancers.
For a quantitative analysis, the Ct of standards with known template concentration is used to generate a
standard curve to which Ct values of unknown samples are compared. The concentration of the unknown
samples is then extrapolated from values from the standard curve. The quantity of amplicons produced in a
PCR reaction is proportional to the prevalence of the targeted sequence; therefore, samples with a higher
template concentration reaches the Ct at earlier PCR cycles than one with a low concentration of the amplified
target. Quantitative q-PCR has high analytical sensitivity for the detection of low mutant allele burden. For
that reason, this method has been widely utilized for monitoring minimal residual disease.
Allele-Specific PCR
Allele-specific PCR (AS-PCR) is a variant of conventional PCR.
The method is based on the principle that Taq polymerase is incapable of catalyzing chain elongation in
the presence of a mismatch between the 3′ end of the primer and the template DNA. Selective amplification
by AS-PCR is achieved by designing a forward primer that matches the mutant sequence at the 3′ end primer.
A second mismatch within the primer can be introduced at the adjacent -1 or -2 position to decrease the
efficiency of mismatched amplification products. This will minimize the chance of amplifying and, therefore,
detecting the wild-type target. AS-PCR is usually performed as two PCR reactions: one employing a forward
primer specific for the mutant sequence, the other using a forward primer specific for the correspondent wild-
type sequence. In this case, a common reverse primer is used for both reactions. Following amplification, the
PCR products are detected by electrophoresis (capillary or agarose gel) or in q-PCR systems. The detection of
adequate PCR product in the wild-type amplification reaction is important to control for adequate specimen
quality and quantity, particularly when the specimen is negative in the mutation-specific PCR reaction.
AS-PCR is particularly useful for the detection of targeted point mutations. Multiplex AS-PCR reactions
can be designed for the simultaneous detection of multiple mutations by including several mutation-specific
primers. The method has high analytical sensitivity and specificity and can be easily deployed in most clinical
laboratories. However, an important limitation is that this approach will not detect mutations other than
those for which specific primers are designed. Therefore, it is utilized for highly recurrent mutations that occur
at specific locations within genes, rather than for the detection of variable mutations that may occur
throughout a gene.
Examples of AS-PCR applications in oncology include the detection of JAK2 V617F and MPL mutations
in myeloproliferative neoplasms (primary myelofibrosis, essential thrombocythemia, and/or polycythemia
vera), the BRAF V600E mutation, and KIT D816V mutations in cases of systemic mastocytosis and in acute
myelogenous leukemia (AML).
Reverse Transcriptase PCR
RT-PCR is utilized for the detection and quantification of RNA transcripts. The first step for all
amplification-based assays that use RNA as a starting material is reverse transcription of RNA into cDNA,
because RNA is not a suitable substrate for Taq polymerase.
In RT-PCR, RNA is isolated and reverse transcribed into cDNA by using a reverse transcriptase enzyme
and one of the following:
(1) random hexamer primers, which anneal randomly to RNA and reverse transcribe all RNA in the cell;
(2) oligo dT primers, which anneal to the polyA tail of mRNA and reverse transcribe only mRNA;
or (3) gene-specific primers that reverse transcribe only the target of interest.
PCR is subsequently performed on the cDNA with forward and reverse primers specific to the gene(s) of
interest. The RT-PCR products may then be analyzed by capillary electrophoresis or in real-time systems as in a
standard PCR reaction.
RT-PCR is commonly used for detecting gene fusions during translocation analysis because breakpoints
frequently occur within the intron of each partner gene and the precise intronic breakpoint locations may be
variable. This variability complicates the design of primers used in DNA-based PCR assays. RT-PCR tests are
advantageous because mature mRNA has intronic sequence spliced out, allowing for simplified primer design
within the affected exon of each partner gene. In this setting, RT-PCR is useful in tests where both
translocation partners are recurrent and only one or a few exons are involved in each partner gene. For
instance, 95% of acute promyelocytic leukemia (APL) cases harbor the reciprocal t(15;17) chromosomal
translocation and these breakpoints always occur within intron 2 of the RARA gene. By contrast, three distinct
chromosome 15 breakpoints are involved, all occurring within the PML gene: intron 6, exon 6, and intron 3.
Because the breakpoints in the two genes are recurrent, most of the reported PML-RARA fusions can be
detected by targeting these three transcript isoforms.
RT-PCR is the method of choice when high sensitivity is required to detect gene translocations. For
example, PML-RARA transcript detection by RT-PCR can detect this fusion transcript down to 1 tumor cell in
the background of 100,000 normal cells.
Detecting low levels of fusion transcript can reveal relapse after consolidation and guide further
treatment. RT-PCR can also be used to quantitate the amount of expression of a gene. One major application
of RT-PCR in this setting includes quantitative detection of BCR-ABL1 fusion transcript for prognostication and
minimal residual disease testing in CML (Fig. 3.4). In this setting, a three log decrease in BCR-ABL1 levels is
associated with an improved outcome.
Fragment Analysis
A fragment analysis is a PCR amplicon-sizing technique that is relevant for the detection of small- to
medium-length–affecting mutations (deletions, insertions, and duplications). This is typically performed by
capillary electrophoresis, which is capable of resolving length mutations from approximately 1 to 500 base
pairs in size.
Fragment analysis represents a practical strategy because it enables comprehensive detection of a
wide variety of possible length mutations and has high analytic sensitivity. Further, it can provide
semiquantitative information regarding the relative amount of mutated alleles. Limitations of this approach
include the inability to objectively quantitate mutant allele burdens, the inability to determine the exact
change in nucleotide sequence, and the inability to detect non–length-affecting mutations such as substitution
mutations.
Examples of fragment analysis applications in oncology include the detection of NPM1 insertion
mutations (Fig. 3.5), EGFR exon 19 deletions, FLT3 internal tandem duplications, and JAK2 exon 12 mutations.
High-Resolution Melting Curve Analysis
A high-resolution melting (HRM) curve analysis is a mutation screening method that allows for the
detection of DNA sequence variations based on specific sequence-related melting profiles of PCR products.
Because the melting property of DNA duplexes is dependent on the biophysical and chemical properties of the
nucleotide sequences, mutant and wild-type DNA sequences can be differentiated from one another based on
their melting characteristics.
An HRM analysis is preceded by a PCR. The reaction employs a pair of gene-specific forward and
reverse primers, template DNA, and a reporter that can either be a double-stranded DNA binding dye or a
fluorescent reporter probe. Following the last cycle of the PCR, the amplification products undergo a cooling
step that generates homoduplexes (double-stranded molecules with perfect complementarity between
alleles) and heteroduplexes (double-stranded molecules with sequence mismatch between alleles) followed
by a heating step that denatures (i.e., melts) the double-stranded products. Heteroduplexes (mutant DNA)
produce a melting profile different from that of wild-type samples (homoduplexes).
In most cases, the reaction is performed in a q-PCR system that allows for an analysis of amplification
and
Vous aimerez peut-être aussi
- Fast Facts: Myelofibrosis: Reviewed by Professor Ruben A. MesaD'EverandFast Facts: Myelofibrosis: Reviewed by Professor Ruben A. MesaPas encore d'évaluation
- Molecularmarkersfor Colorectalcancer: Moriah Wright,, Jenifer S. Beaty,, Charles A. TernentDocument19 pagesMolecularmarkersfor Colorectalcancer: Moriah Wright,, Jenifer S. Beaty,, Charles A. TernentFernando Castro EchavezPas encore d'évaluation
- 10 5772@54652 PDFDocument32 pages10 5772@54652 PDFhollPas encore d'évaluation
- Minimal Residual Disease in Acute Lymphoblastic Leukemia: Dario CampanaDocument6 pagesMinimal Residual Disease in Acute Lymphoblastic Leukemia: Dario CampanaJAIR_valencia3395Pas encore d'évaluation
- 2012 - Intra-Tumour Heterogeneity - A Looking Glass For CancerDocument12 pages2012 - Intra-Tumour Heterogeneity - A Looking Glass For CancerFreddy A. ManayayPas encore d'évaluation
- Tugas DR KamalDocument7 pagesTugas DR KamalZarin SafanahPas encore d'évaluation
- Acute Myeloid Leukemia: Advances in Diagnosis and Classi CationDocument9 pagesAcute Myeloid Leukemia: Advances in Diagnosis and Classi CationTay SalinasPas encore d'évaluation
- 225 VannucchiDocument10 pages225 VannucchiAwais ArshadPas encore d'évaluation
- Chapitre de Livre BoutainaDocument37 pagesChapitre de Livre BoutainaBoutaina AddoumPas encore d'évaluation
- Chronic Lymphocytic Leukemia: A Clinical and Molecular Heterogenous DiseaseDocument14 pagesChronic Lymphocytic Leukemia: A Clinical and Molecular Heterogenous DiseaseCallisthenisLeventisPas encore d'évaluation
- Cancer Total BothDocument33 pagesCancer Total BothUsman AshrafPas encore d'évaluation
- Prognostic Factors in Acute Myeloid Leukaemia 4: Bob LoèwenbergDocument11 pagesPrognostic Factors in Acute Myeloid Leukaemia 4: Bob LoèwenbergStephania SandovalPas encore d'évaluation
- 2020 Genetic Markers in Lung Cancer Diagnosis A ReviewDocument22 pages2020 Genetic Markers in Lung Cancer Diagnosis A Review191933033196Pas encore d'évaluation
- MR 25Document9 pagesMR 25Riko JumattullahPas encore d'évaluation
- Deros A 2015Document10 pagesDeros A 2015rakaPas encore d'évaluation
- Cytogenetic Analysis in The Diagnosis of Acute Leukemia: Sverre Heim, FelixDocument9 pagesCytogenetic Analysis in The Diagnosis of Acute Leukemia: Sverre Heim, FelixEnas KharbotlyPas encore d'évaluation
- Neuroblastoma: Biological Insights Into A Clinical Enigma: Garrett M. BrodeurDocument14 pagesNeuroblastoma: Biological Insights Into A Clinical Enigma: Garrett M. BrodeurMouris DwiputraPas encore d'évaluation
- Ijms 19 03492Document33 pagesIjms 19 03492Qaasim DramatPas encore d'évaluation
- Brock Et Al Liquid Biopsy 2015Document11 pagesBrock Et Al Liquid Biopsy 2015Jean BiverPas encore d'évaluation
- Genetic Alterations and DNA Repair in Human Carcinogenesis: Kathleen Dixon, Elizabeth KoprasDocument8 pagesGenetic Alterations and DNA Repair in Human Carcinogenesis: Kathleen Dixon, Elizabeth Kopras1205lorenaPas encore d'évaluation
- Biomarkers For The Lung Cancer Diagnosis and Their Advances in ProteomicsDocument11 pagesBiomarkers For The Lung Cancer Diagnosis and Their Advances in ProteomicsAndi Harmawati NPas encore d'évaluation
- ADVANCE Molecular Techniques and Histopathology Applications ShortenedDocument5 pagesADVANCE Molecular Techniques and Histopathology Applications ShortenedDale TelgenhoffPas encore d'évaluation
- Chapter 1 - The Cancer GenomeDocument35 pagesChapter 1 - The Cancer GenomeCynthia LopesPas encore d'évaluation
- Unidad VDocument75 pagesUnidad VpablenquePas encore d'évaluation
- Tmp9a87 TMPDocument3 pagesTmp9a87 TMPFrontiersPas encore d'évaluation
- CM7144 Saoirsemorrin 19333910Document51 pagesCM7144 Saoirsemorrin 19333910Saoirse MorrinPas encore d'évaluation
- Plasma Cell Myeloma Literature Review and Case StudyDocument6 pagesPlasma Cell Myeloma Literature Review and Case StudyafdtygyhkPas encore d'évaluation
- Pancreatic CancerDocument13 pagesPancreatic CancerFA MonterPas encore d'évaluation
- Molecular Oncology: Directory of ServicesDocument31 pagesMolecular Oncology: Directory of Servicesruby_kakkar9796100% (2)
- Glioblastoma Stem-Like Cells Approaches For Isolation and CharacterizationDocument19 pagesGlioblastoma Stem-Like Cells Approaches For Isolation and CharacterizationMariano PerezPas encore d'évaluation
- New England Journal Medicine: The ofDocument16 pagesNew England Journal Medicine: The ofMauricio FemeníaPas encore d'évaluation
- Classification of Adult Acute Myeloid LeukemiaDocument24 pagesClassification of Adult Acute Myeloid LeukemiaSusianna RismandaPas encore d'évaluation
- Clinical Implications of BRAF Mutation Test in Colorectal CancerDocument8 pagesClinical Implications of BRAF Mutation Test in Colorectal CancerMohammed AladhraeiPas encore d'évaluation
- Understanding Familial and Non-Familial Renal Cell CancerDocument10 pagesUnderstanding Familial and Non-Familial Renal Cell CancerVanroPas encore d'évaluation
- Molecular Biology As A Tool To Treat CancerDocument8 pagesMolecular Biology As A Tool To Treat CancerFreelance EduPas encore d'évaluation
- Haake Et Al 2017 CancerDocument10 pagesHaake Et Al 2017 Cancerdiego peinadoPas encore d'évaluation
- Giskeødegård Et Al. - Unknown - NMR Based Metabolomics of Biofluids in CancerDocument28 pagesGiskeødegård Et Al. - Unknown - NMR Based Metabolomics of Biofluids in Canceryannick brunatoPas encore d'évaluation
- 1) Explain The Risk Factors For CLL, Including The Relevance of Monoclonal B Cell Lymphocytosis (MBL)Document6 pages1) Explain The Risk Factors For CLL, Including The Relevance of Monoclonal B Cell Lymphocytosis (MBL)Fahim MahmudPas encore d'évaluation
- Pediatricneuro-Oncology: Fatema MalbariDocument17 pagesPediatricneuro-Oncology: Fatema MalbariRandy UlloaPas encore d'évaluation
- The Risk Factors For CLL, Including The Relevance of Monoclonal B Cell Lymphocytosis (MBL)Document7 pagesThe Risk Factors For CLL, Including The Relevance of Monoclonal B Cell Lymphocytosis (MBL)Fahim MahmudPas encore d'évaluation
- Roylance 1997Document8 pagesRoylance 1997barti koksPas encore d'évaluation
- DownloadDocument12 pagesDownloadhasan nazzalPas encore d'évaluation
- JCR 23 007Document13 pagesJCR 23 007yakalapunagarajuPas encore d'évaluation
- Colorectal Cancer PredispositionDocument3 pagesColorectal Cancer PredispositionPete FlintPas encore d'évaluation
- Myeloid Malignancies: Mutations, Models and Management: Review Open AccessDocument15 pagesMyeloid Malignancies: Mutations, Models and Management: Review Open AccessZikry Aulia100% (1)
- 531 (1999) T. R. Golub: Science Et AlDocument8 pages531 (1999) T. R. Golub: Science Et AlBair PuigPas encore d'évaluation
- 1 s2.0 S0304383523000083 MainDocument9 pages1 s2.0 S0304383523000083 MainMericia Guadalupe Sandoval ChavezPas encore d'évaluation
- Tumor ProgressionDocument7 pagesTumor ProgressionmineresearchPas encore d'évaluation
- Cell Cycle and Cancer - Unit I IV/IDocument6 pagesCell Cycle and Cancer - Unit I IV/IManoj GourojuPas encore d'évaluation
- Pi Is 0923753420360798Document16 pagesPi Is 0923753420360798ekanovicaPas encore d'évaluation
- 10 Use of Flow Cytometry in Clonal Plasma Cell DisordersDocument7 pages10 Use of Flow Cytometry in Clonal Plasma Cell DisorderscandiddreamsPas encore d'évaluation
- 5472.can 16 1578Document42 pages5472.can 16 1578肖茹雪Pas encore d'évaluation
- Cancer Res 1990 Wainscoat 1355 60Document7 pagesCancer Res 1990 Wainscoat 1355 60Shahab Ud DinPas encore d'évaluation
- Acute Myeloid Leukemia A Concise ReviewDocument17 pagesAcute Myeloid Leukemia A Concise Reviewberlianza activiraPas encore d'évaluation
- Adult Acute Lymphoblastic Leukemia: Concepts and StrategiesDocument12 pagesAdult Acute Lymphoblastic Leukemia: Concepts and StrategiesdrravesPas encore d'évaluation
- 13 ChemoDocument40 pages13 ChemoSweet SunPas encore d'évaluation
- AlsnlasnfDocument13 pagesAlsnlasnfDina A. ŠabićPas encore d'évaluation
- Acute Promyelocytic Leukemia Part-1Document8 pagesAcute Promyelocytic Leukemia Part-1Francisco Ignacio Reiser ValverdePas encore d'évaluation
- Pages 4 7Document4 pagesPages 4 7andreas_251650Pas encore d'évaluation
- Review Article WHO Classification of Tumors of Hematopoietic and Lymphoid Tissues 4th EditionDocument8 pagesReview Article WHO Classification of Tumors of Hematopoietic and Lymphoid Tissues 4th EditionCat NhanPas encore d'évaluation
- AbemaciclibDocument2 pagesAbemaciclibIna SimachePas encore d'évaluation
- Hallmarks of Cancer An OrganizingDocument4 pagesHallmarks of Cancer An OrganizingIna SimachePas encore d'évaluation
- Alkylating AgentsDocument3 pagesAlkylating AgentsIna SimachePas encore d'évaluation
- Autoimmune Disorders Associated To Type 1Document10 pagesAutoimmune Disorders Associated To Type 1Ina SimachePas encore d'évaluation
- Unified Power Quality Conditioner (Upqc) With Pi and Hysteresis Controller For Power Quality Improvement in Distribution SystemsDocument7 pagesUnified Power Quality Conditioner (Upqc) With Pi and Hysteresis Controller For Power Quality Improvement in Distribution SystemsKANNAN MANIPas encore d'évaluation
- KAHOOT - Assignment 4.1 Lesson PlanDocument3 pagesKAHOOT - Assignment 4.1 Lesson PlanJan ZimmermannPas encore d'évaluation
- 1stQ Week5Document3 pages1stQ Week5Jesse QuingaPas encore d'évaluation
- Hercules Industries Inc. v. Secretary of Labor (1992)Document1 pageHercules Industries Inc. v. Secretary of Labor (1992)Vianca MiguelPas encore d'évaluation
- Julie Jacko - Professor of Healthcare InformaticsDocument1 pageJulie Jacko - Professor of Healthcare InformaticsjuliejackoPas encore d'évaluation
- SEx 3Document33 pagesSEx 3Amir Madani100% (4)
- Windows SCADA Disturbance Capture: User's GuideDocument23 pagesWindows SCADA Disturbance Capture: User's GuideANDREA LILIANA BAUTISTA ACEVEDOPas encore d'évaluation
- Islamic Meditation (Full) PDFDocument10 pagesIslamic Meditation (Full) PDFIslamicfaith Introspection0% (1)
- Lorraln - Corson, Solutions Manual For Electromagnetism - Principles and Applications PDFDocument93 pagesLorraln - Corson, Solutions Manual For Electromagnetism - Principles and Applications PDFc. sorasPas encore d'évaluation
- Barangay Sindalan v. CA G.R. No. 150640, March 22, 2007Document17 pagesBarangay Sindalan v. CA G.R. No. 150640, March 22, 2007FD BalitaPas encore d'évaluation
- Your Free Buyer Persona TemplateDocument8 pagesYour Free Buyer Persona Templateel_nakdjoPas encore d'évaluation
- The Preparedness of The Data Center College of The Philippines To The Flexible Learning Amidst Covid-19 PandemicDocument16 pagesThe Preparedness of The Data Center College of The Philippines To The Flexible Learning Amidst Covid-19 PandemicInternational Journal of Innovative Science and Research TechnologyPas encore d'évaluation
- Accounting 110: Acc110Document19 pagesAccounting 110: Acc110ahoffm05100% (1)
- Hilti Product Technical GuideDocument16 pagesHilti Product Technical Guidegabox707Pas encore d'évaluation
- Kormos - Csizer Language Learning 2008Document29 pagesKormos - Csizer Language Learning 2008Anonymous rDHWR8eBPas encore d'évaluation
- Research ProposalDocument18 pagesResearch ProposalIsmaelPas encore d'évaluation
- Lite Touch. Completo PDFDocument206 pagesLite Touch. Completo PDFkerlystefaniaPas encore d'évaluation
- The Training Toolbox: Forced Reps - The Real Strength SenseiDocument7 pagesThe Training Toolbox: Forced Reps - The Real Strength SenseiSean DrewPas encore d'évaluation
- Name Numerology Calculator - Chaldean Name Number PredictionsDocument2 pagesName Numerology Calculator - Chaldean Name Number Predictionsarunamurugesan7Pas encore d'évaluation
- Focus Group DiscussionDocument13 pagesFocus Group DiscussionSumon ChowdhuryPas encore d'évaluation
- Progressivism Sweeps The NationDocument4 pagesProgressivism Sweeps The NationZach WedelPas encore d'évaluation
- The 5 RS:: A New Teaching Approach To Encourage Slowmations (Student-Generated Animations) of Science ConceptsDocument7 pagesThe 5 RS:: A New Teaching Approach To Encourage Slowmations (Student-Generated Animations) of Science Conceptsnmsharif66Pas encore d'évaluation
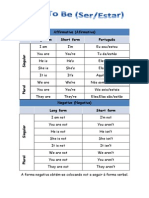
- Affirmative (Afirmativa) Long Form Short Form PortuguêsDocument3 pagesAffirmative (Afirmativa) Long Form Short Form PortuguêsAnitaYangPas encore d'évaluation
- Blunders and How To Avoid Them Dunnington PDFDocument147 pagesBlunders and How To Avoid Them Dunnington PDFrajveer404100% (2)
- Algebra. Equations. Solving Quadratic Equations B PDFDocument1 pageAlgebra. Equations. Solving Quadratic Equations B PDFRoberto CastroPas encore d'évaluation
- Clearing Negative SpiritsDocument6 pagesClearing Negative SpiritsmehorseblessedPas encore d'évaluation
- Nursing Care Plan: Pt.'s Data Nursing Diagnosis GoalsDocument1 pageNursing Care Plan: Pt.'s Data Nursing Diagnosis GoalsKiran Ali100% (3)
- Fansubbers The Case of The Czech Republic and PolandDocument9 pagesFansubbers The Case of The Czech Republic and Polandmusafir24Pas encore d'évaluation
- Leg Res Cases 4Document97 pagesLeg Res Cases 4acheron_pPas encore d'évaluation
- Sino-Japanese Haikai PDFDocument240 pagesSino-Japanese Haikai PDFAlina Diana BratosinPas encore d'évaluation
- 10% Human: How Your Body's Microbes Hold the Key to Health and HappinessD'Everand10% Human: How Your Body's Microbes Hold the Key to Health and HappinessÉvaluation : 4 sur 5 étoiles4/5 (33)
- Masterminds: Genius, DNA, and the Quest to Rewrite LifeD'EverandMasterminds: Genius, DNA, and the Quest to Rewrite LifePas encore d'évaluation
- Why We Die: The New Science of Aging and the Quest for ImmortalityD'EverandWhy We Die: The New Science of Aging and the Quest for ImmortalityÉvaluation : 4.5 sur 5 étoiles4.5/5 (6)
- When the Body Says No by Gabor Maté: Key Takeaways, Summary & AnalysisD'EverandWhen the Body Says No by Gabor Maté: Key Takeaways, Summary & AnalysisÉvaluation : 3.5 sur 5 étoiles3.5/5 (2)
- Tales from Both Sides of the Brain: A Life in NeuroscienceD'EverandTales from Both Sides of the Brain: A Life in NeuroscienceÉvaluation : 3 sur 5 étoiles3/5 (18)
- The Rise and Fall of the Dinosaurs: A New History of a Lost WorldD'EverandThe Rise and Fall of the Dinosaurs: A New History of a Lost WorldÉvaluation : 4 sur 5 étoiles4/5 (597)
- Buddha's Brain: The Practical Neuroscience of Happiness, Love & WisdomD'EverandBuddha's Brain: The Practical Neuroscience of Happiness, Love & WisdomÉvaluation : 4 sur 5 étoiles4/5 (216)
- Return of the God Hypothesis: Three Scientific Discoveries That Reveal the Mind Behind the UniverseD'EverandReturn of the God Hypothesis: Three Scientific Discoveries That Reveal the Mind Behind the UniverseÉvaluation : 4.5 sur 5 étoiles4.5/5 (52)
- A Brief History of Intelligence: Evolution, AI, and the Five Breakthroughs That Made Our BrainsD'EverandA Brief History of Intelligence: Evolution, AI, and the Five Breakthroughs That Made Our BrainsÉvaluation : 4.5 sur 5 étoiles4.5/5 (6)
- The Molecule of More: How a Single Chemical in Your Brain Drives Love, Sex, and Creativity--and Will Determine the Fate of the Human RaceD'EverandThe Molecule of More: How a Single Chemical in Your Brain Drives Love, Sex, and Creativity--and Will Determine the Fate of the Human RaceÉvaluation : 4.5 sur 5 étoiles4.5/5 (517)
- Gut: the new and revised Sunday Times bestsellerD'EverandGut: the new and revised Sunday Times bestsellerÉvaluation : 4 sur 5 étoiles4/5 (393)
- Seven and a Half Lessons About the BrainD'EverandSeven and a Half Lessons About the BrainÉvaluation : 4 sur 5 étoiles4/5 (111)
- Undeniable: How Biology Confirms Our Intuition That Life Is DesignedD'EverandUndeniable: How Biology Confirms Our Intuition That Life Is DesignedÉvaluation : 4 sur 5 étoiles4/5 (11)
- All That Remains: A Renowned Forensic Scientist on Death, Mortality, and Solving CrimesD'EverandAll That Remains: A Renowned Forensic Scientist on Death, Mortality, and Solving CrimesÉvaluation : 4.5 sur 5 étoiles4.5/5 (397)
- The Ancestor's Tale: A Pilgrimage to the Dawn of EvolutionD'EverandThe Ancestor's Tale: A Pilgrimage to the Dawn of EvolutionÉvaluation : 4 sur 5 étoiles4/5 (812)
- Who's in Charge?: Free Will and the Science of the BrainD'EverandWho's in Charge?: Free Will and the Science of the BrainÉvaluation : 4 sur 5 étoiles4/5 (65)
- Good Without God: What a Billion Nonreligious People Do BelieveD'EverandGood Without God: What a Billion Nonreligious People Do BelieveÉvaluation : 4 sur 5 étoiles4/5 (66)
- Moral Tribes: Emotion, Reason, and the Gap Between Us and ThemD'EverandMoral Tribes: Emotion, Reason, and the Gap Between Us and ThemÉvaluation : 4.5 sur 5 étoiles4.5/5 (116)
- Remnants of Ancient Life: The New Science of Old FossilsD'EverandRemnants of Ancient Life: The New Science of Old FossilsÉvaluation : 3 sur 5 étoiles3/5 (3)
- The Lives of Bees: The Untold Story of the Honey Bee in the WildD'EverandThe Lives of Bees: The Untold Story of the Honey Bee in the WildÉvaluation : 4.5 sur 5 étoiles4.5/5 (44)
- Darwin's Doubt: The Explosive Origin of Animal Life and the Case for Intelligent DesignD'EverandDarwin's Doubt: The Explosive Origin of Animal Life and the Case for Intelligent DesignÉvaluation : 4 sur 5 étoiles4/5 (19)
- The Consciousness Instinct: Unraveling the Mystery of How the Brain Makes the MindD'EverandThe Consciousness Instinct: Unraveling the Mystery of How the Brain Makes the MindÉvaluation : 4.5 sur 5 étoiles4.5/5 (93)
- Human: The Science Behind What Makes Your Brain UniqueD'EverandHuman: The Science Behind What Makes Your Brain UniqueÉvaluation : 3.5 sur 5 étoiles3.5/5 (38)
- The Other Side of Normal: How Biology Is Providing the Clues to Unlock the Secrets of Normal and Abnormal BehaviorD'EverandThe Other Side of Normal: How Biology Is Providing the Clues to Unlock the Secrets of Normal and Abnormal BehaviorPas encore d'évaluation